Biochemical and structural characterization of a novel family of cystathionine beta-synthase domain proteins fused to a Zn ribbon-like domain
Proudfoot, M., Sanders, S.A., Singer, A., Zhang, R., Brown, G., Binkowski, A., Xu, L., Lukin, J.A., Murzin, A.G., Joachimiak, A., Arrowsmith, C.H., Edwards, A.M., Savchenko, A.V., Yakunin, A.F.(2008) J Mol Biol 375: 301-315
- PubMed: 18021800
- DOI: https://doi.org/10.1016/j.jmb.2007.10.060
- Primary Citation of Related Structures:
1PVM, 2QH1 - PubMed Abstract:
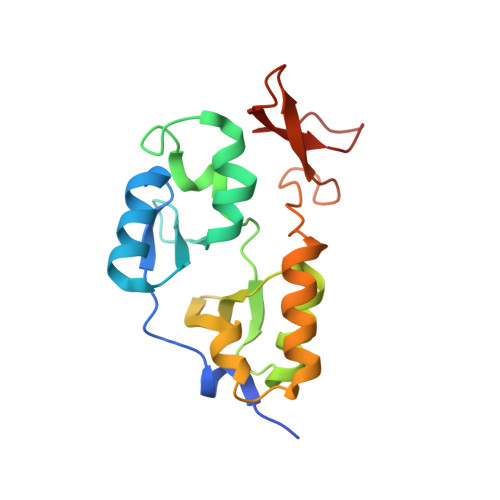
We have identified a novel family of proteins, in which the N-terminal cystathionine beta-synthase (CBS) domain is fused to the C-terminal Zn ribbon domain. Four proteins were overexpressed in Escherichia coli and purified: TA0289 from Thermoplasma acidophilum, TV1335 from Thermoplasma volcanium, PF1953 from Pyrococcus furiosus, and PH0267 from Pyrococcus horikoshii. The purified proteins had a red/purple color in solution and an absorption spectrum typical of rubredoxins (Rds). Metal analysis of purified proteins revealed the presence of several metals, with iron and zinc being the most abundant metals (2-67% of iron and 12-74% of zinc). Crystal structures of both mercury- and iron-bound TA0289 (1.5-2.0 A resolution) revealed a dimeric protein whose intersubunit contacts are formed exclusively by the alpha-helices of two cystathionine beta-synthase subdomains, whereas the C-terminal domain has a classical Zn ribbon planar architecture. All proteins were reversibly reduced by chemical reductants (ascorbate or dithionite) or by the general Rd reductase NorW from E. coli in the presence of NADH. Reduced TA0289 was found to be capable of transferring electrons to cytochrome C from horse heart. Likewise, the purified Zn ribbon protein KTI11 from Saccharomyces cerevisiae had a purple color in solution and an Rd-like absorption spectrum, contained both iron and zinc, and was reduced by the Rd reductase NorW from E. coli. Thus, recombinant Zn ribbon domains from archaea and yeast demonstrate an Rd-like electron carrier activity in vitro. We suggest that, in vivo, some Zn ribbon domains might also bind iron and therefore possess an electron carrier activity, adding another physiological role to this large family of important proteins.
Organizational Affiliation:
Banting and Best Department of Medical Research, University of Toronto, 112 College Street, Room 72, Toronto, ON, Canada.