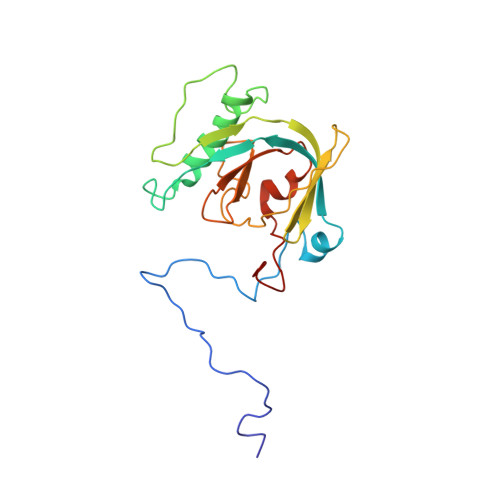
Coupling of Folding and Binding in the PTB Domain of the Signaling Protein Shc
Farooq, A., Zeng, L., Yan, K.S., Ravichandran, K.S., Zhou, M.-M.(2003) Structure 11: 905-913
- PubMed: 12906822
- DOI: https://doi.org/10.1016/s0969-2126(03)00134-5
- Primary Citation of Related Structures:
1N3H, 1OY2 - PubMed Abstract:
The notion that certain proteins lack intrinsic globular structure under physiological conditions and that the attainment of fully folded structure only occurs upon the binding of target molecules has been recently gaining popularity. We report here the solution structure of the PTB domain of the signaling protein Shc in the free form. Comparison of this structure with that of the complex form, obtained previously with a phosphopeptide ligand, reveals that the Shc PTB domain is structurally disordered in the free form, particularly around the regions constituting the peptide binding pocket. The binding of the ligand appears to reorganize this pocket through local folding events triggering a conformational switch between the free and the complex forms.
Organizational Affiliation:
Structural Biology Program, Department of Physiology and Biophysics, Mount Sinai School of Medicine, New York University, New York, NY 10029, USA. amjad.farooq@mssm.edu