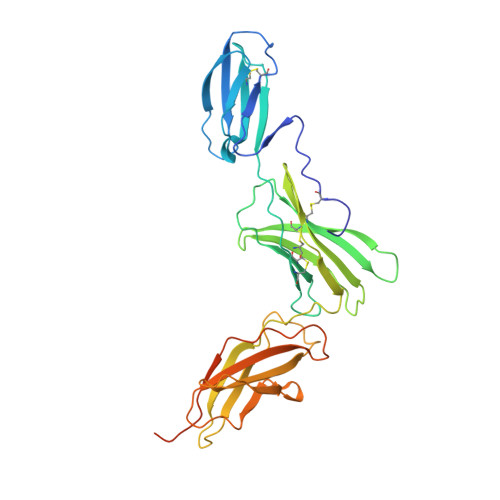
Structure of the extracellular domains of the human interleukin-6 receptor alpha-chain.
Varghese, J.N., Moritz, R.L., Lou, M.-Z., van Donkelaar, A., Ji, H., Ivancic, N., Branson, K.M., Hall, N.E., Simpson, R.J.(2002) Proc Natl Acad Sci U S A 99: 15959-15964
- PubMed: 12461182
- DOI: https://doi.org/10.1073/pnas.232432399
- Primary Citation of Related Structures:
1N26 - PubMed Abstract:
Dysregulated production of IL-6 and its receptor (IL-6R) are implicated in the pathogenesis of multiple myeloma, autoimmune diseases and prostate cancer. The IL-6R complex comprises two molecules each of IL-6, IL-6R, and the signaling molecule, gp130. Here, we report the x-ray structure (2.4 A) of the IL-6R ectodomains. The N-terminal strand of the Ig-like domain (D(1)) is disulfide-bonded to domain D(2), and domains D(2) and D(3), the cytokine-binding domain, are structurally similar to known cytokine-binding domains. The head-to-tail packing of two closely associated IL-6R molecules observed in the crystal may be representative of the configuration of the physiological dimer of IL-6R and provides new insight into the architecture of the IL-6R complex.
Organizational Affiliation:
Biomolecular Research Institute and Commonwealth Scientific and Industrial Research Organization Health Sciences and Nutrition, 343 Royal Parade, Parkville 3052, Australia.