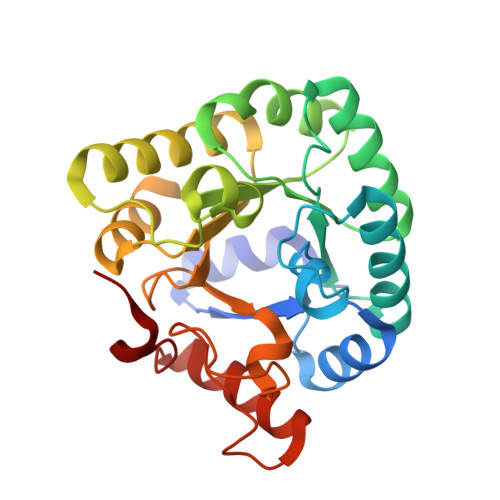
Structure of E. coli Ketopantoate Hydroxymethyl Transferase Complexed with Ketopantoate and Mg(2+), Solved by Locating 160 Selenomethionine Sites.
von Delft, F., Inoue, T., Saldanha, S.A., Ottenhof, H.H., Schmitzberger, F., Birch, L.M., Dhanaraj, V., Witty, M., Smith, A.G., Blundell, T.L., Abell, C.(2003) Structure 11: 985-996
- PubMed: 12906829
- DOI: https://doi.org/10.1016/s0969-2126(03)00158-8
- Primary Citation of Related Structures:
1M3U - PubMed Abstract:
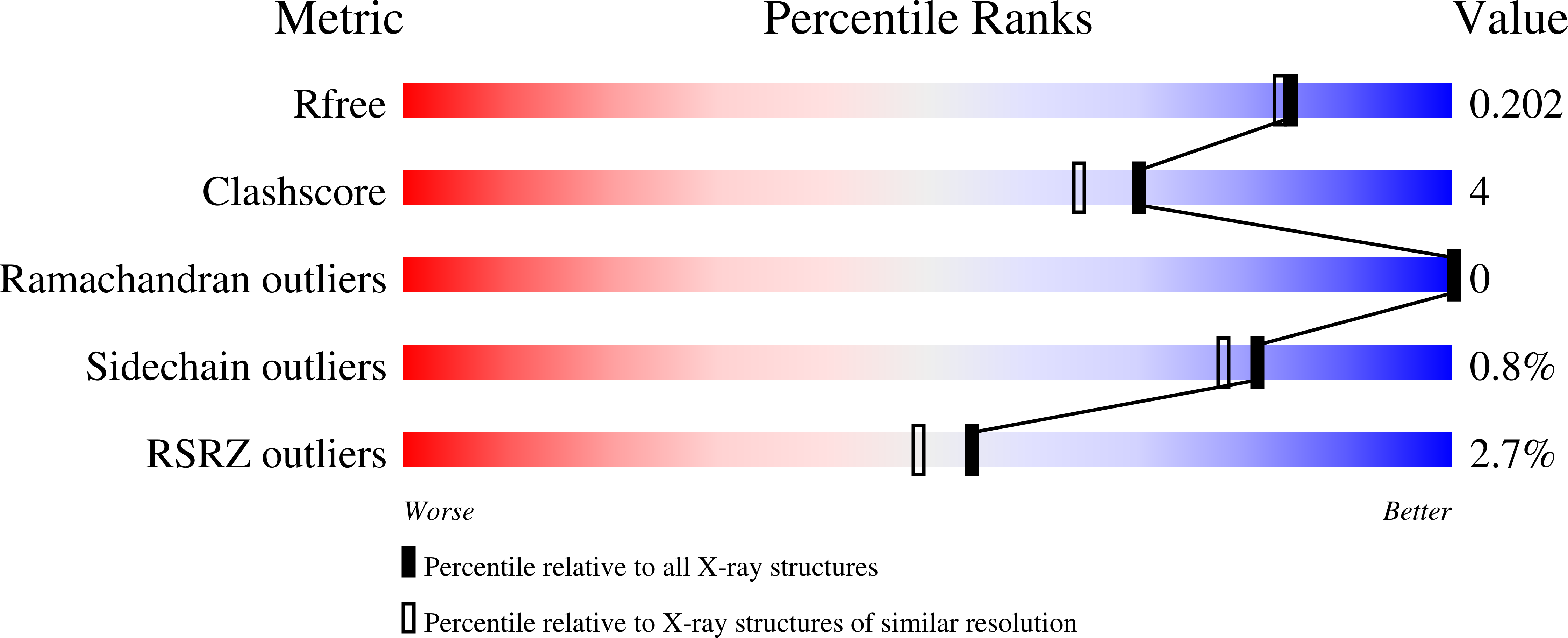
We report the crystal structure of E. coli ketopantoate hydroxymethyltransferase (KPHMT) at 1.9 A resolution, in complex with its product, ketopantoate. KPHMT catalyzes the first step in the biosynthesis of pantothenate (vitamin B(5)), the precursor of coenzyme A and the acyl carrier protein cofactor. The structure of the decameric enzyme was solved by multiwavelength anomalous dispersion to locate 160 selenomethionine sites and phase 560 kDa of protein, making it the largest structure solved by this approach. KPHMT adopts the (betaalpha)(8) barrel fold and is a member of the phosphoenolpyruvate/pyruvate superfamily. The active site contains a ketopantoate bidentately coordinated to Mg(2+). Similar binding is likely for the substrate, alpha-ketoisovalerate, orienting the C3 for deprotonation.
Organizational Affiliation:
Department of Biochemistry, 80 Tennis Court Road, Cambridge CB2 1GA, United Kingdom.