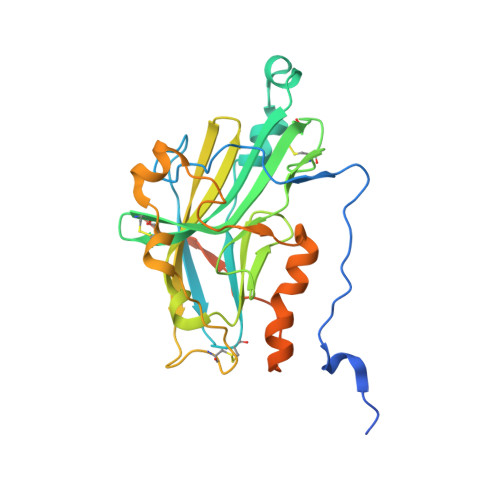
Herpes simplex virus glycoprotein D bound to the human receptor HveA.
Carfi, A., Willis, S.H., Whitbeck, J.C., Krummenacher, C., Cohen, G.H., Eisenberg, R.J., Wiley, D.C.(2001) Mol Cell 8: 169-179
- PubMed: 11511370
- DOI: https://doi.org/10.1016/s1097-2765(01)00298-2
- Primary Citation of Related Structures:
1JMA, 1L2G - PubMed Abstract:
Herpes simplex virus (HSV) infection requires binding of the viral envelope glycoprotein D (gD) to cell surface receptors. We report the X-ray structures of a soluble, truncated ectodomain of gD both alone and in complex with the ectodomain of its cellular receptor HveA. Two bound anions suggest possible binding sites for another gD receptor, a 3-O-sulfonated heparan sulfate. Unexpectedly, the structures reveal a V-like immunoglobulin (Ig) fold at the core of gD that is closely related to cellular adhesion molecules and flanked by large N- and C-terminal extensions. The receptor binding segment of gD, an N-terminal hairpin, appears conformationally flexible, suggesting that a conformational change accompanying binding might be part of the viral entry mechanism.
Organizational Affiliation:
Department of Medicine, Children's Hospital, Howard Hughes Medical Institute, 320 Longwood Avenue, Boston, MA 02115, USA.