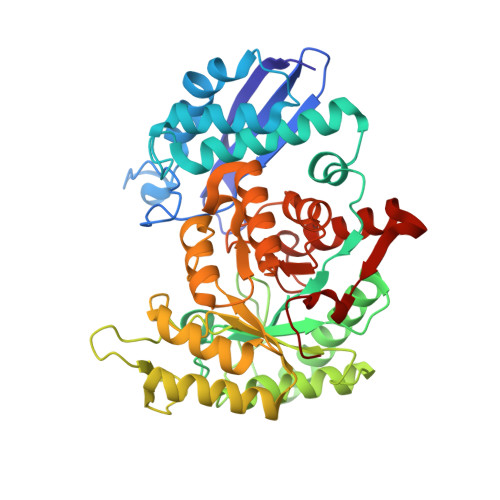
Crystal Structure of Enterococcus hirae Enolase at 2.8 A Resolution
Hosaka, T., Meguro, T., Yamato, I., Shirakihara, Y.(2003) J Biochem 133: 817-823
- PubMed: 12869539
- DOI: https://doi.org/10.1093/jb/mvg104
- Primary Citation of Related Structures:
1IYX - PubMed Abstract:
We report the crystal structure of an enolase from Enterococcus hirae, which is the first report of a structure determination among gram-positive bacteria. We isolated the enolase gene and determined the base sequence. The amino acid sequence deduced from the DNA sequence suggests that this enolase is composed of 431 amino acids. The amino acid sequence is very similar to those of enolases from eukaryotic and prokaryotic organisms, being 65% and 50% identical to enolases from Escherichia coli and yeast, respectively. The enolase prepared from E. hirae lysate yielded crystals containing one dimer per asymmetric unit. X-ray diffraction patterns were obtained at 2.8 A resolution on a SPring-8 synchrotron radiation source. Crystals belong to space group I4 with unit cell dimensions of a = b = 153.5 A, c = 90.7 A. The E. hirae, yeast, E. coli and lobster enolase structures are very similar. The E. hirae enolase takes an "Open" conformation. The regions in the structure that differ most from other enolases are loops L4 (132-140) and L3 (244-265). Considering the positions of these loops relative to the active site, they seem to have no direct involvement in function. Our findings show that the three dimensional structure of an important enzyme in the glycolytic pathway is evolutionarily conserved among eukaryotes and prokaryotes, including gram-positive bacteria.
Organizational Affiliation:
Department of Biological Science and Technology, Tokyo University of Science, 2641 Yamazaki, Noda-shi, Chiba 278-8510.