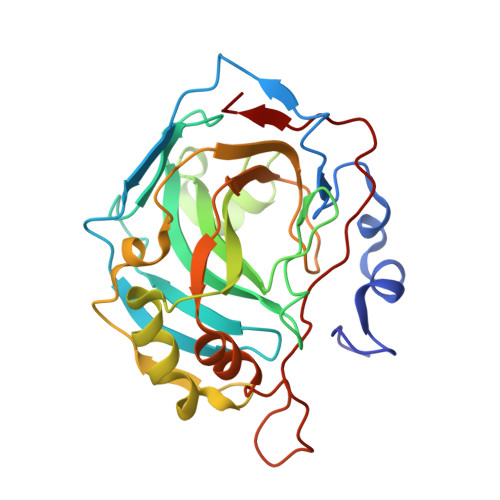
Structural aspects of isozyme selectivity in the binding of inhibitors to carbonic anhydrases II and IV.
Kim, C.Y., Whittington, D.A., Chang, J.S., Liao, J., May, J.A., Christianson, D.W.(2002) J Med Chem 45: 888-893
- PubMed: 11831900
- DOI: https://doi.org/10.1021/jm010163d
- Primary Citation of Related Structures:
1I8Z, 1I90, 1I91 - PubMed Abstract:
Carbonic anhydrase inhibitors are effective in lowering intraocular pressure, the primary indication of glaucoma. Human carbonic anhydrase II, and possibly carbonic anhydrase IV (CAII and CAIV, respectively), help regulate fluid secretion into the anterior chamber of the eye. Because inhibitors currently formulated as drugs to treat glaucoma were designed to target CAII, an understanding of the structural basis of CAII-CAIV discrimination by inhibitors would be useful for probing the role of each isozyme in the etiology of the disease. Here, we report the X-ray crystal structures of three novel thieno[3,2-e]-1,2-thiazine-6-sulfonamides complexed with CAII and the computationally predicted structures of the same compounds complexed with CAIV. All three compounds bind with similar affinity to CAII, but they bind with up to 100-fold lower affinities to CAIV. Comparisons of experimentally determined structures of CAII-inhibitor complexes and computationally predicted structures of CAIV-inhibitor complexes allow us to rationalize these affinity trends and outline molecular features that may contribute to high-affinity inhibitor binding to CAIV. This study demonstrates how experimental structure determination methods and computational structure prediction methods can be used together to answer questions that cannot be answered by either method alone.
Organizational Affiliation:
Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, PA 19104-6323, USA.