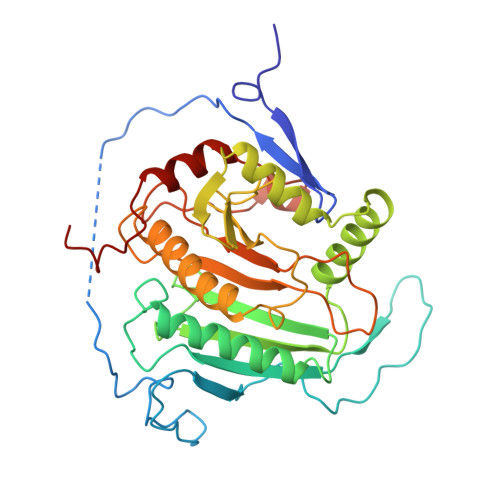
Three-dimensional structure of galactose-1-phosphate uridylyltransferase from Escherichia coli at 1.8 A resolution.
Wedekind, J.E., Frey, P.A., Rayment, I.(1995) Biochemistry 34: 11049-11061
- PubMed: 7669762
- DOI: https://doi.org/10.1021/bi00035a010
- Primary Citation of Related Structures:
1HXP - PubMed Abstract:
Galactose-1-phosphate uridylyltransferase catalyzes the reversible transfer of the uridine 5'-monophosphoryl moiety of UDP-glucose to the phosphate group of galactose 1-phosphate to form UDP-galactose. This enzyme participates in the Leloir pathway of galactose metabolism, and its absence is the primary cause of the potentially lethal disease galactosemia. The three-dimensional structure of the dimeric enzyme from Escherichia coli complexed with uridine 5'-diphosphate is reported here. The structure was solved by multiple isomorphous replacement and electron density modification techniques and has been refined to 1.8 A resolution. Enzyme subunits consist of a single domain with the topology of a "half-barrel". The barrel staves are formed by nine strands of antiparallel beta-sheet. The barrel axis is approximately parallel to the local dyad that relates each subunit. Two amphipathic helices fill the half-barrel sequestering its hydrophobic interior. An iron atom resides on the outside of the barrel, centered in the subunit interface. Intrasubunit coordination to iron resembles a distorted square pyramid formed by the equatorial ligation of two histidines and a bidentate carboxylate group and a single axial histidine. The subunit interface is stabilized by this coordination and is further characterized by the formation of two intermolecular "mini-sheets" distinct from the strands of the half-barrel. Loops that connect the mini-sheet strands contribute to the formation of the active site, which resides on the external surface of the barrel rim. Loops of the barrel strands are tethered together by a structural zinc atom that orients the local fold in a manner essential for catalysis. In one of the latter loops, S gamma of a cysteine is modified by beta-mercaptoethanol, which prevents the alpha-phosphorus of the nucleotide from access to the nucleophile His166. This conformation does not appear to perturb the interactions to the uracil and ribose moieties as mediated through the side chains of Leu54, Ohe75, Asn77, Asp78, Phe79, and Val108. Several of the latter residues have been implicated in human galactosemia. The present structure explains the deleterious effects of many of those mutations.
Organizational Affiliation:
Institute for Enzyme Research, Graduate School, University of Wisconsin-Madison 53705, USA.