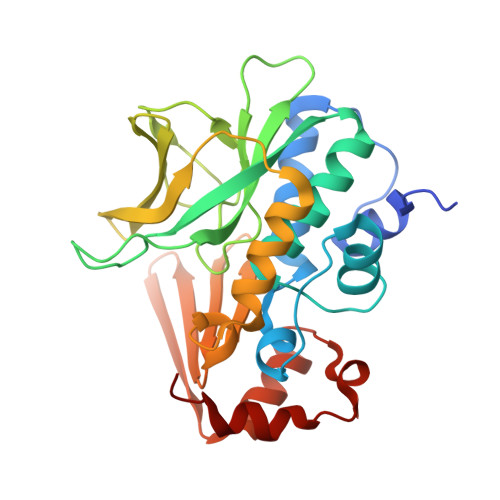
The Structure of Arylamine N-Acetyltransferase from Mycobacterium Smegmatis-an Enzyme which Inactivates the Anti-Tubercular Drug, Isoniazid
Sandy, J., Mushtaq, A., Kawamura, A., Sinclair, J., Sim, E., Noble, M.(2002) J Mol Biol 318: 1071
- PubMed: 12054803
- DOI: https://doi.org/10.1016/S0022-2836(02)00141-9
- Primary Citation of Related Structures:
1GX3 - PubMed Abstract:
Arylamine N-acetyltransferases which acetylate and inactivate isoniazid, an anti-tubercular drug, are found in mycobacteria including Mycobacterium smegmatis and Mycobacterium tuberculosis. We have solved the structure of arylamine N-acetyltransferase from M. smegmatis at a resolution of 1.7 A as a model for the highly homologous NAT from M. tuberculosis. The fold closely resembles that of NAT from Salmonella typhimurium, with a common catalytic triad and domain structure that is similar to certain cysteine proteases. The detailed geometry of the catalytic triad is typical of enzymes which use primary alcohols or thiols as activated nucleophiles. Thermal mobility and structural variations identify parts of NAT which might undergo conformational changes during catalysis. Sequence conservation among eubacterial NATs is restricted to structural residues of the protein core, as well as the active site and a hinge that connects the first two domains of the NAT structure. The structure of M. smegmatis NAT provides a template for modelling the structure of the M. tuberculosis enzyme and for structure-based ligand design as an approach to designing anti-TB drugs.
Organizational Affiliation:
Department of Pharmacology, University of Oxford, Mansfield Road, OX1 3QT, UK.