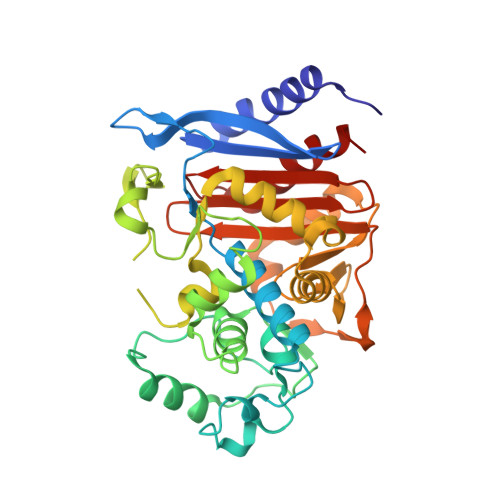
Structure of the extended-spectrum class C beta-lactamase of Enterobacter cloacae GC1, a natural mutant with a tandem tripeptide insertion.
Crichlow, G.V., Kuzin, A.P., Nukaga, M., Mayama, K., Sawai, T., Knox, J.R.(1999) Biochemistry 38: 10256-10261
- PubMed: 10441119
- DOI: https://doi.org/10.1021/bi9908787
- Primary Citation of Related Structures:
1GCE - PubMed Abstract:
A class C beta-lactamase from a clinical isolate of Enterobacter cloacae strain GC1 with improved hydrolytic activity for oxyimino beta-lactam antibiotics has been analyzed by X-ray crystallography to 1.8 A resolution. Relative to the wild-type P99 beta-lactamase, this natural mutant contains a highly unique tandem repeat Ala211-Val212-Arg213 [Nugaka et al. (1995) J. Biol. Chem. 270, 5729-5735]. The 39.4 kDa chromosomal beta-lactamase crystallizes from poly(ethylene glycol) 8000 in potassium phosphate in space group P2(1)2(1)2 with cell dimensions a = 78.0 A, b = 69.5 A, and c = 63.1 A. The crystal structure was solved by the molecular replacement method, and the model has been refined to an R-factor of 0.20 for all nonzero data from 8 to 1.8 A. Deviations of model bonds and angles from ideal values are 0.008 A and 1.4 degrees, respectively. Overlay of alpha-carbon atoms in the GC1 and P99 beta-lactamases results in an rms deviation of 0.6 A. Largest deviations occur in a loop containing Gln120 and in the Omega loop region (200-218) where the three residues 213-215 are disordered. Possibly as a result of this disorder, the width of the opening to the substrate binding cavity, as measured from the 318-324 beta-strand to two loops containing Gln120 and Tyr150 on the other side, is 0.6-1.4 A wider than in P99. It is suggested that conformational flexibility in the expanded Omega loop, and its influence on adjacent protein structure, may facilitate hydrolysis of oxyimino beta-lactams by making the acyl intermediate more open to attack by water. Nevertheless, backbone atoms in core catalytic site residues Ser64, Lys67, Tyr150, Asn152, Lys318, and Ser321 deviate only 0.4 A (rmsd) from atoms in P99. A rotation of a potential catalytic base, Tyr150, relative to P99 at pH 8, is consistent with the requirement for a lower than normal pK(a) for this residue.
Organizational Affiliation:
Department of Molecular and Cell Biology, The University of Connecticut, Storrs 06269-3125, USA.