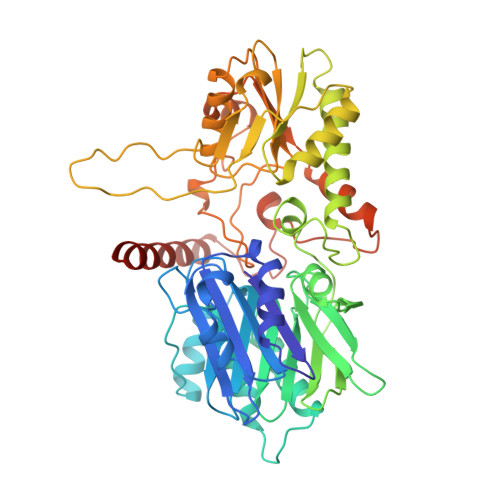
Structure and function of the glutamine phosphoribosylpyrophosphate amidotransferase glutamine site and communication with the phosphoribosylpyrophosphate site.
Kim, J.H., Krahn, J.M., Tomchick, D.R., Smith, J.L., Zalkin, H.(1996) J Biol Chem 271: 15549-15557
- PubMed: 8663035
- DOI: https://doi.org/10.1074/jbc.271.26.15549
- Primary Citation of Related Structures:
1ECG - PubMed Abstract:
Glutamine phosphoribosylpyrophosphate (PRPP) amidotransferase from Escherichia coli exhibits a basal PRPP-independent glutaminase activity having a kcat/Km that is 0.3% of fully active enzyme. Binding of PRPP activates the enzyme by a structural change that lowers the Km for glutamine 100-fold and couples glutamine hydrolysis to synthesis of 5-phosphoribosylamine. By analysis of the x-ray structure of the glutamine site containing bound 6-diazo-5-oxonorleucine, a glutamine affinity analog, and by site-directed mutagenesis we have identified residues important for glutamine binding, catalysis, and coupling with PRPP. Tyr74 is a key residue in the coupling between the sites for glutamine in the NH2-terminal domain and PRPP in the COOH-terminal domain. Arg73 and Asp127 have roles in glutamine binding. The x-ray structure indicates that there are no amino acid side chains sufficiently close to Cys1 to participate as a proton acceptor in formation of the thiolate needed for nucleophilic attack on the carboxamide of glutamine, nor as a general acid for amide nitrogen transfer. Based on the x-ray model of the glutamine site and analysis of a mutant enzyme we propose that the free NH2 terminus of Cys1 functions as the proton acceptor and donor. The results indicate that the side chain of Asn101 and the backbone nitrogen of Gly102 function to stabilize a tetrahedral oxyanion resulting from attack of Cys1 on the glutamine carboxamide. Cys1, Arg73, Asn101, Gly102, and Asp127 are conserved in the NH2-terminal domain of a subfamily of amidotransferases that includes asparagine synthetase, glucosamine 6-phosphate synthase, and glutamate synthase, implying a common function in the four enzymes. Tyr74, on the other hand, is conserved only in glutamine PRPP amidotransferase sequences consistent with a specific role in interdomain coupling. The catalytic framework of key glutamine site residues supports the assignment of glutamine PRPP amidotransferase to a recently described Ntn (NH2-terminal nucleophile) hydrolase family of enzymes.
Organizational Affiliation:
Department of Biochemistry, Purdue University, West Lafayette, Indiana 47907-1153, USA.