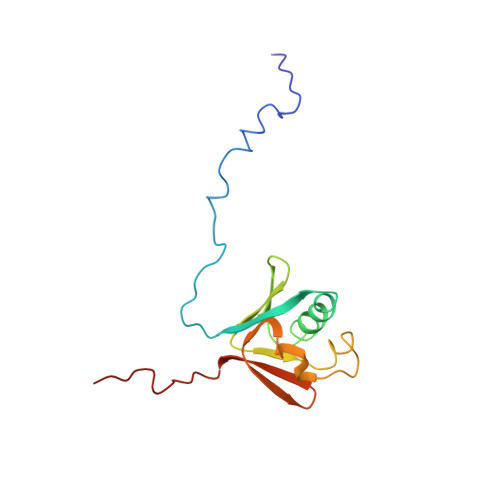
Structure of the soluble methane monooxygenase regulatory protein B.
Walters, K.J., Gassner, G.T., Lippard, S.J., Wagner, G.(1999) Proc Natl Acad Sci U S A 96: 7877-7882
- PubMed: 10393915
- DOI: https://doi.org/10.1073/pnas.96.14.7877
- Primary Citation of Related Structures:
1CKV - PubMed Abstract:
The soluble methane monooxygenase (sMMO; EC 1.14.13.25) from the pseudothermophile Methylococcus capsulatus (Bath) is a three-component enzyme system that catalyzes the selective oxidation of methane to methanol. We have used NMR spectroscopy to produce a highly refined structure of MMOB, the 16-kDa regulatory protein of this system. This structure has a unique and intricate fold containing seven beta-strands forming two beta-sheets oriented perpendicular to each other and bridged by three alpha-helices. The rate and efficiency of the methane hydroxylation by sMMO depend on dynamic binding interactions of the hydroxylase with the reductase and regulatory protein components during catalysis. We have monitored by NMR the binding of MMOB to the hydroxylase in the presence and absence of the reductase. The results of these studies provide structural insight into how the regulatory protein interacts with the hydroxylase.
Organizational Affiliation:
Committee on Higher Degrees in Biophysics, Harvard University, Cambridge, MA 02138, USA.