Atomic Structure of the Trypsin-Aeruginosin 98-B Complex
Sandler, B., Murakami, M., Clardy, J.(1998) J Am Chem Soc 120: 595-596
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
(1998) J Am Chem Soc 120: 595-596
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| TRYPSIN | 223 | Bos taurus | Mutation(s): 0 EC: 3.4.21.4 | 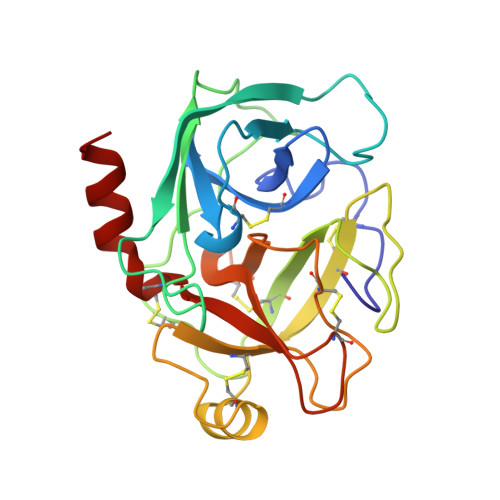 | |
UniProt | |||||
Find proteins for P00760 (Bos taurus) Explore P00760 Go to UniProtKB: P00760 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00760 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| AERUGINOSIN 98-B | 4 | Microcystis aeruginosa NIES-98 | Mutation(s): 0 | 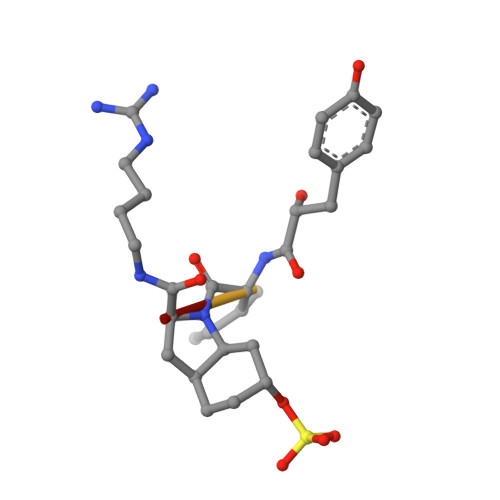 | |
Sequence AnnotationsExpand | |||||
| |||||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| XPR Query on XPR | B | L-PEPTIDE LINKING | C9 H15 N O6 S |  | PRO |
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_000556 Query on PRD_000556 | B | AERUGINOSIN 98-B | Oligopeptide / Thrombin inhibitor, Trypsin inhibitor |  | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 48.3 | α = 90 |
| b = 48.3 | β = 90 |
| c = 145.2 | γ = 120 |
| Software Name | Purpose |
|---|---|
| DENZO | data reduction |
| SCALEPACK | data scaling |
| X-PLOR | model building |
| X-PLOR | refinement |
| X-PLOR | phasing |