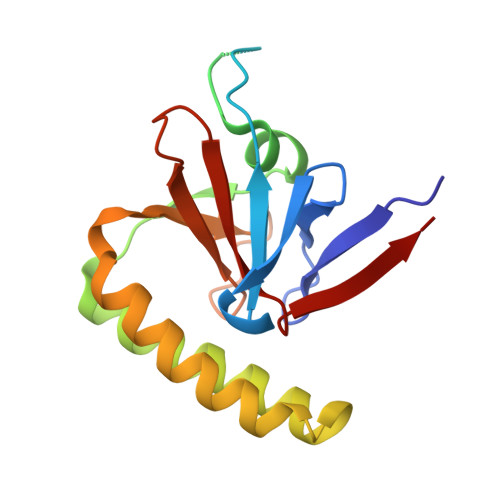
TCTP protects from apoptotic cell death by antagonizing bax function
Susini, L., Besse, S., Duflaut, D., Lespagnol, A., Beekman, C., Fiucci, G., Atkinson, A.R., Busso, D., Poussin, P., Marine, J.C., Martinou, J.C., Cavarelli, J., Moras, D., Amson, R., Telerman, A.(2008) Cell Death Differ 15: 1211-1220
- PubMed: 18274553
- DOI: https://doi.org/10.1038/cdd.2008.18
- Primary Citation of Related Structures:
1YZ1 - PubMed Abstract:
Translationally controlled tumor protein (TCTP) is a potential target for cancer therapy. It functions as a growth regulating protein implicated in the TSC1-TSC2 -mTOR pathway or a guanine nucleotide dissociation inhibitor for the elongation factors EF1A and EF1Bbeta. Accumulating evidence indicates that TCTP also functions as an antiapoptotic protein, through a hitherto unknown mechanism. In keeping with this, we show here that loss of tctp expression in mice leads to increased spontaneous apoptosis during embryogenesis and causes lethality between E6.5 and E9.5. To gain further mechanistic insights into this apoptotic function, we solved and refined the crystal structure of human TCTP at 2.0 A resolution. We found a structural similarity between the H2-H3 helices of TCTP and the H5-H6 helices of Bax, which have been previously implicated in regulating the mitochondrial membrane permeability during apoptosis. By site-directed mutagenesis we establish the relevance of the H2-H3 helices in TCTP's antiapoptotic function. Finally, we show that TCTP antagonizes apoptosis by inserting into the mitochondrial membrane and inhibiting Bax dimerization. Together, these data therefore further confirm the antiapoptotic role of TCTP in vivo and provide new mechanistic insights into this key function of TCTP.
Organizational Affiliation:
Molecular Engines Laboratories, 20 rue Bouvier, Paris, France.