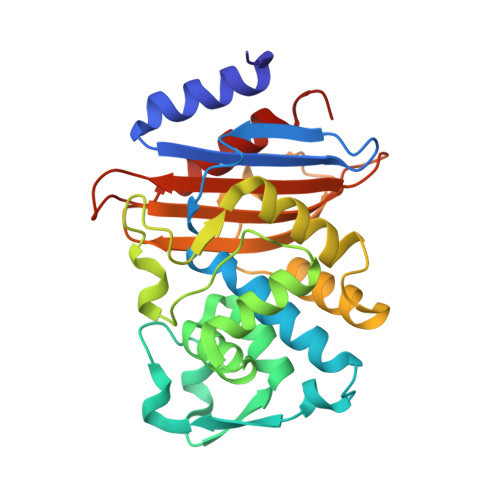
Atomic Resolution Structures of CTX-M beta-Lactamases: Extended Spectrum Activities from Increased Mobility and Decreased Stability.
Chen, Y., Delmas, J., Sirot, J., Shoichet, B., Bonnet, R.(2005) J Mol Biol 348: 349-362
- PubMed: 15811373
- DOI: https://doi.org/10.1016/j.jmb.2005.02.010
- Primary Citation of Related Structures:
1YLJ, 1YLP, 1YLT, 1YLW - PubMed Abstract:
Extended spectrum beta-lactamases (ESBLs) confer bacterial resistance to third-generation cephalosporins, such as cefotaxime and ceftazidime, increasing hospital mortality rates. Whereas these antibiotics are almost impervious to classic beta-lactamases, such as TEM-1, ESBLs have one to four orders greater activity against them. The origins of this activity have been widely studied for the TEM and SHV-type ESBLs, but have received less attention for the CTX-M beta-lactamases, an emerging family that is now the dominant ESBL in several regions. To understand how CTX-M beta-lactamases achieve their remarkable activity, biophysical and structural studies were undertaken. Using reversible, two-state thermal denaturation, it was found that as these enzymes evolve a broader substrate range, they sacrifice stability. Thus, the mutant enzyme CTX-M-16 is eightfold more active against ceftazidime than the pseudo-wild-type CTX-M-14 but is 1.9 kcal/mol less stable. This is consistent with a "stability-activity tradeoff," similar to that observed in the evolution of other resistance enzymes. To investigate the structural basis of enzyme activity and stability, the structures of four CTX-M enzymes were determined by X-ray crystallography. The structures of CTX-M-14, CTX-M-27, CTX-M-9 and CTX-M-16 were determined to 1.10 Angstroms, 1.20 Angstroms, 0.98 Angstroms and 1.74 Angstroms resolution, respectively. The enzyme active sites resemble those of the narrow-spectrum TEM-1 and SHV-1, and not the enlarged sites typical of ESBL mutants such as TEM-52 and TEM-64. Instead, point substitutions leading to specific interactions may be responsible for the improved activity against ceftazidime and cefotaxime, consistent with observations first made for the related Toho-1 enzyme. The broadened substrate range of CTX-M-16 may result from coupled defects in the enzyme's B3 strand, which lines the active site. Substitutions Val231-->Ala and Asp240-->Gly, which convert CTX-M-14 into CTX-M-16, occur at either end of this strand. These defects appear to increase the mobility of B3 based on anisotropic B-factor analyses at ultrahigh resolution, consistent with stability loss and activity gain. The unusually high resolution of these structures that makes such analyses possible also makes them good templates for inhibitor discovery.
Organizational Affiliation:
Department of Pharmaceutical Chemistry, University of California, San Francisco, Genentech Hall, 600 16th Street, San Francisco, CA 94143-2240, USA.