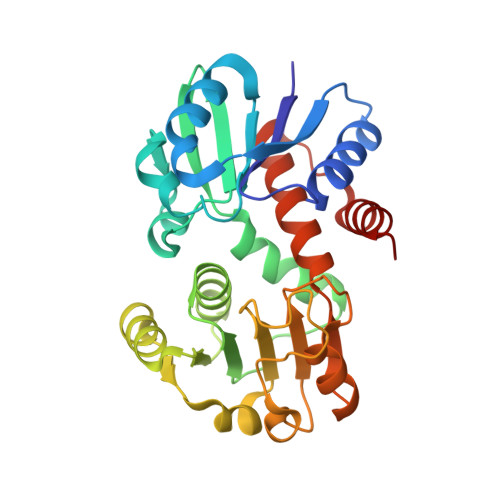
Crystal Structures of Shikimate Dehydrogenase AroE from Thermus thermophilus HB8 and its Cofactor and Substrate Complexes: Insights into the Enzymatic Mechanism
Bagautdinov, B., Kunishima, N.(2007) J Mol Biol 373: 424-438
- PubMed: 17825835
- DOI: https://doi.org/10.1016/j.jmb.2007.08.017
- Primary Citation of Related Structures:
1WXD, 2CY0, 2D5C, 2EV9 - PubMed Abstract:
Shikimate dehydrogenase (EC 1.1.1.25) catalyses the fourth step of the shikimate pathway which is required for the synthesis of the aromatic amino acids and other aromatic compounds in bacteria, microbial eukaryotes, and plants. The crystal structures of the shikimate dehydrogenase AroE from Thermus thermophilus HB8 in its ligand-free form, binary complexes with cofactor NADP+ or substrate shikimate, and the ternary complex with both NADP(H) and shikimate were determined by X-ray diffraction method at atomic resolutions. The crystals are nearly isomorphous with the asymmetric unit containing a dimer, each subunit of which has a bi-domain structure of compact alpha/beta sandwich folds. The two subunits of the enzyme display asymmetry in the crystals due to different relative orientations between the N- and C-terminal domains resulting in a slightly different closure of the interdomain clefts. NADP(H) is bound to the more closed form only. This closed conformation with apparent higher affinity to the cofactor is also observed in the unliganded crystal form, indicating that the NADP(H) binding to TtAroE may follow the selection mode where the cofactor binds to the subunit that happens to be in the closed conformation in solution. Crystal structures of the closed subunits with and without NADP(H) show no significant structural difference, suggesting that the cofactor binding to the closed subunit corresponds to the lock-and-key model in TtAroE. On the other hand, shikimate binds to both open and closed subunit conformers of both apo and NADP(H)-liganded holo enzyme forms. The ternary complex TtAroE:NADP(H):shikimate allows unambiguous visualization of the SDH permitting elucidation of the roles of conserved residues Lys64 and Asp100 in the hydride ion transfer between NADP(H) and shikimate.
Organizational Affiliation:
Advanced Protein Crystallography Research Group, RIKEN SPring-8 Center, Harima Institute, 1-1-1 Kouto, Sayo-cho, Sayo-gun, Hyogo 679-5148, Japan.