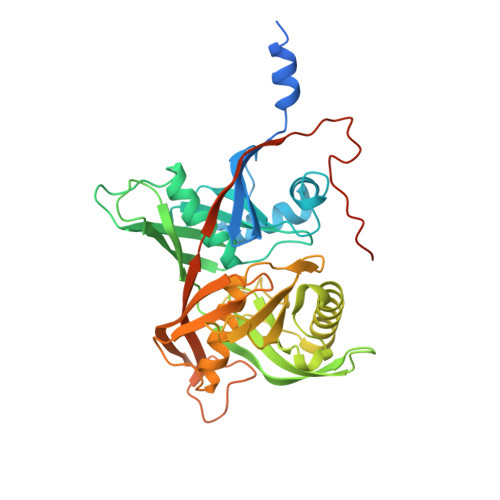
Crystal structure, catalytic mechanism, and mitogenic properties of Trypanosoma cruzi proline racemase.
Buschiazzo, A., Goytia, M., Schaeffer, F., Degrave, W., Shepard, W., Gregoire, C., Chamond, N., Cosson, A., Berneman, A., Coatnoan, N., Alzari, P.M., Minoprio, P.(2006) Proc Natl Acad Sci U S A 103: 1705-1710
- PubMed: 16446443
- DOI: https://doi.org/10.1073/pnas.0509010103
- Primary Citation of Related Structures:
1W61, 1W62 - PubMed Abstract:
Amino acid racemases catalyze the stereoinversion of the chiral C alpha to produce the d-enantiomers that participate in biological processes, such as cell wall construction in prokaryotes. Within this large protein family, bacterial proline racemases have been extensively studied as a model of enzymes acting with a pyridoxal-phosphate-independent mechanism. Here we report the crystal structure of the proline racemase from the human parasite Trypanosoma cruzi (TcPRACA), a secreted enzyme that triggers host B cell polyclonal activation, which prevents specific humoral immune responses and is crucial for parasite evasion and fate. The enzyme is a homodimer, with each monomer folded in two symmetric alpha/beta subunits separated by a deep crevice. The structure of TcPRACA in complex with a transition-state analog, pyrrole-2-carboxylic acid, reveals the presence of one reaction center per monomer, with two Cys residues optimally located to perform acid/base catalysis through a carbanion stabilization mechanism. Mutation of the catalytic Cys residues abolishes the enzymatic activity but preserves the mitogenic properties of the protein. In contrast, inhibitor binding promotes the closure of the interdomain crevice and completely abrogates B cell proliferation, suggesting that the mitogenic properties of TcPRACA depend on the exposure of transient epitopes in the ligand-free enzyme.
Organizational Affiliation:
Unité de Biochimie Structurale, Centre National de la Recherche Scientifique/Institut Pasteur, 25 Rue du Dr. Roux, F-75724 Paris, France.