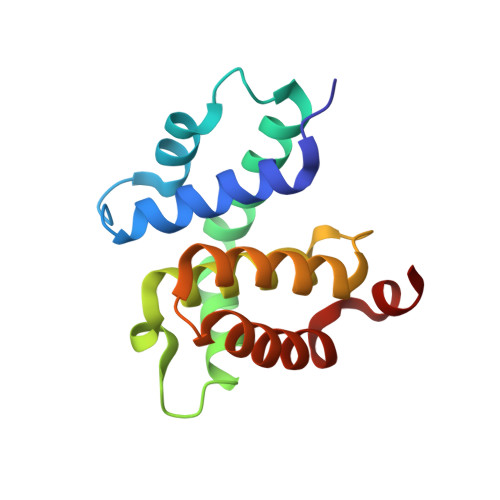
Crystal structures of the antitermination factor NusB from Thermotoga maritima and implications for RNA binding
Bonin, I., Robelek, R., Benecke, H., Urlaub, H., Bacher, A., Richter, G., Wahl, M.C.(2004) Biochem J 383: 419-428
- PubMed: 15279620
- DOI: https://doi.org/10.1042/BJ20040889
- Primary Citation of Related Structures:
1TZT, 1TZU, 1TZV, 1TZW, 1TZX - PubMed Abstract:
NusB is a prokaryotic transcription factor involved in antitermination processes, during which it interacts with the boxA portion of the mRNA nut site. Previous studies have shown that NusB exhibits an all-helical fold, and that the protein from Escherichia coli forms monomers, while Mycobacterium tuberculosis NusB is a dimer. The functional significance of NusB dimerization is unknown. We have determined five crystal structures of NusB from Thermotoga maritima. In three crystal forms the protein appeared monomeric, whereas the two other crystal forms contained assemblies, which resembled the M. tuberculosis dimers. In solution, T. maritima NusB could be cross-linked as dimers, but it migrated as a monomer in gel-filtration analyses, suggesting a monomer/dimer equilibrium with a preference for the monomer. Binding to boxA-like RNA sequences could be detected by gel-shift analyses and UV-induced cross-linking. An N-terminal arginine-rich sequence is a probable RNA binding site of the protein, exhibiting aromatic residues as potential stacking partners for the RNA bases. Anions located in various structures support the assignment of this RNA binding site. The proposed RNA binding region is hidden in the subunit interface of dimeric NusB proteins, such as NusB from M. tuberculosis, suggesting that such dimers have to undergo a considerable conformational change or dissociate for engagement with RNA. Therefore, in certain organisms, dimerization may be employed to package NusB in an inactive form until recruitment into antitermination complexes.
Organizational Affiliation:
Max-Planck Institut für Biochemie, Abteilung Strukturforschung, Am Klopferspitz 18a, D-82152 Martinsried, Germany.