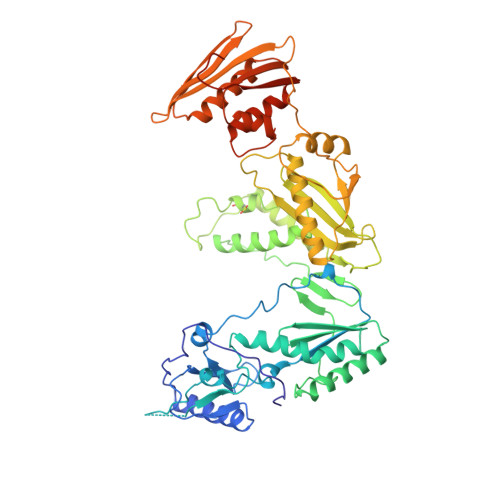
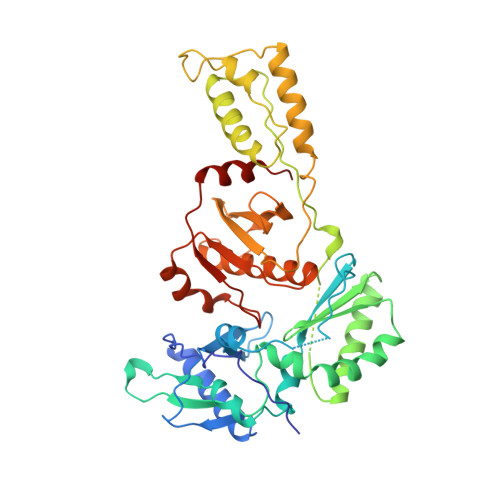
Crystal structures of HIV-1 reverse transcriptase in complex with carboxanilide derivatives.
Ren, J., Esnouf, R.M., Hopkins, A.L., Warren, J., Balzarini, J., Stuart, D.I., Stammers, D.K.(1998) Biochemistry 37: 14394-14403
- PubMed: 9772165
- DOI: https://doi.org/10.1021/bi981309m
- Primary Citation of Related Structures:
1RT4, 1RT5, 1RT6, 1RT7 - PubMed Abstract:
The carboxanilides are nonnucleoside inhibitors (NNIs) of HIV-1 reverse transcriptase (RT), of potential clinical importance. The compounds differ in potency and in their retention of potency in the face of drug resistance mutations. Whereas UC-84, the prototype compound, only weakly inhibits many RTs bearing single point resistance mutations, inhibition by UC-781 is little affected. It has been proposed that UC-38 and UC-781 may form quaternary complexes with RT at a site other than the known binding pocket of other NNIs. X-ray crystal structures of four HIV-1 RT-carboxanilide complexes (UC-10, UC-38, UC-84, and UC-781) reported here reveal that all four inhibitors bind in the usual NNI site, forming binary 1:1 complexes with RT in the absence of substrates with the amide/thioamide bond in cis conformations. For all four complexes the anilide rings of the inhibitors overlap aromatic rings of many other NNIs bound to RT. In contrast, the second rings of UC-10, UC-84, and UC-781 do not bind in equivalent positions to those of other "two-ring" NNIs such as alpha-APA or HEPT derivatives. The binding modes most closely resemble that of the structurally dissimilar NNI, Cl-TIBO, with a common hydrogen bond between each carboxanilide NH- group and the main-chain carbonyl oxygen of Lys101. The binding modes differ slightly between the UC-10/UC-781 and UC-38/UC-84 pairs of compounds, apparently related to the shorter isopropylmethanoyl substituents of the anilide rings of UC-38/UC-84, which draws these rings closer to residues Tyr181 and Tyr188. This in turn explains the differences in the effect of mutated residues on the binding of these compounds.
Organizational Affiliation:
Laboratory of Molecular Biophysics, Oxford, UK.