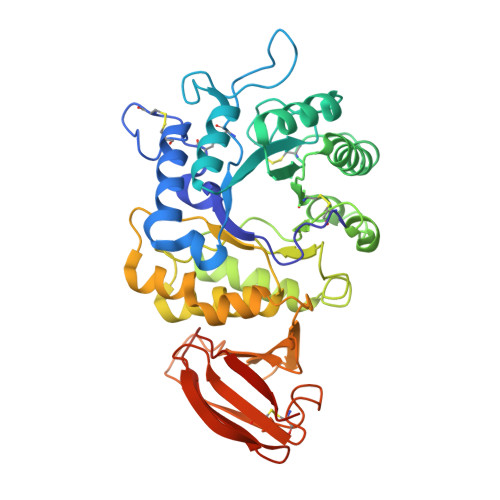
The molecular defect leading to Fabry disease: structure of human alpha-galactosidase
Garman, S.C., Garboczi, D.N.(2004) J Mol Biol 337: 319-335
- PubMed: 15003450
- DOI: https://doi.org/10.1016/j.jmb.2004.01.035
- Primary Citation of Related Structures:
1R46, 1R47 - PubMed Abstract:
Fabry disease is an X-linked lysosomal storage disease afflicting 1 in 40,000 males with chronic pain, vascular degeneration, cardiac impairment, and other symptoms. Deficiency in the lysosomal enzyme alpha-galactosidase (alpha-GAL) causes an accumulation of its substrate, which ultimately leads to Fabry disease symptoms. Here, we present the structure of the human alpha-GAL glycoprotein determined by X-ray crystallography. The structure is a homodimer with each monomer containing a (beta/alpha)8 domain with the active site and an antiparallel beta domain. N-linked carbohydrate appears at six sites in the glycoprotein dimer, revealing the basis for lysosomal transport via the mannose-6-phosphate receptor. To understand how the enzyme cleaves galactose from glycoproteins and glycolipids, we also determined the structure of the complex of alpha-GAL with its catalytic product. The catalytic mechanism of the enzyme is revealed by the location of two aspartic acid residues (D170 and D231), which act as a nucleophile and an acid/base, respectively. As a point mutation in alpha-GAL can lead to Fabry disease, we have catalogued and plotted the locations of 245 missense and nonsense mutations in the three-dimensional structure. The structure of human alpha-GAL brings Fabry disease into the realm of molecular diseases, where insights into the structural basis of the disease phenotypes might help guide the clinical treatment of patients.
Organizational Affiliation:
Structural Biology Section, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Twinbrook II, 12441 Parklawn Drive, Rockville, MD 20852, USA. sgarman@niaid.nih.gov