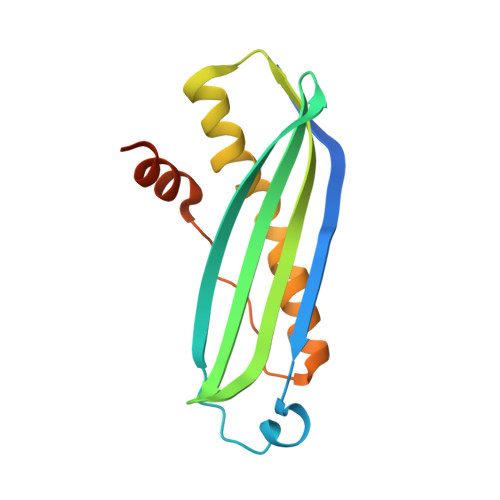
Crystal structure of SecB from Escherichia coli
Dekker, C., de Kruijff, B., Gros, P.(2003) J Struct Biol 144: 313-319
- PubMed: 14643199
- DOI: https://doi.org/10.1016/j.jsb.2003.09.012
- Primary Citation of Related Structures:
1QYN - PubMed Abstract:
The chaperone SecB from Escherichia coli is primarily involved in passing precursor proteins into the Sec system via specific interactions with SecA. The crystal structure of SecB from E. coli has been solved to 2.35 A resolution. The structure shows flexibility in the crossover loop and the helix-connecting loop, regions that have been implicated to be part of the SecB substrate-binding site. Moreover conformational variability of Trp36 is observed as well as different loop conformations for the different monomers. Based on this, we speculate that SecB can regulate the access or extent of its hydrophobic substrate-binding site, by modulating the conformation of the crossover loop and the helix-connecting loop. The structure also clearly explains why the tetrameric equilibrium is shifted towards the dimeric state in the mutant SecBCys76Tyr. The buried cysteine residue is crucial for tight packing, and mutations are likely to disrupt the tetramer formation but not the dimer formation.
Organizational Affiliation:
National Institute for Medical Research, The Ridgeway, Mill Hill, London, England, NW7 1AA, UK. cdekker@mimr.mrc.ac.uk