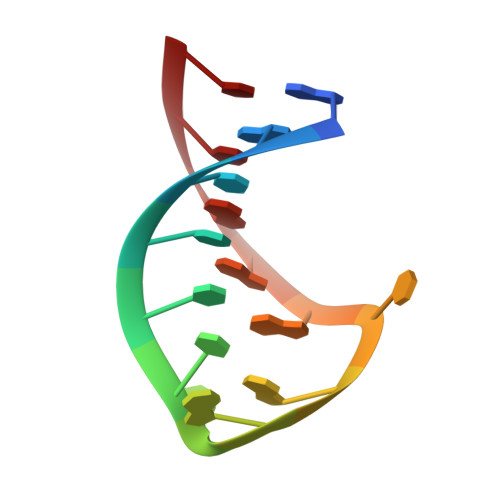
YNMG tetraloop formation by a dyskeratosis congenita mutation in human telomerase RNA.
Theimer, C.A., Finger, L.D., Feigon, J.(2003) RNA 9: 1446-1455
- PubMed: 14624001
- DOI: https://doi.org/10.1261/rna.5152303
- Primary Citation of Related Structures:
1Q75 - PubMed Abstract:
Autosomal dominant dyskeratosis congenita (DKC) has been linked to mutations in the RNA component of telomerase, the ribonucleoprotein responsible for telomere maintenance. Recent studies have investigated the role of the GC (107-108) --> AG mutation in the conserved P3 helix in the pseudoknot domain of human telomerase RNA. The mutation was found to significantly destabilize the pseudoknot conformation, resulting in a shift in the thermodynamic equilibrium to favor formation of a P2b hairpin intermediate. In the wild-type sequence, the hairpin intermediate was found to form a novel sequence of pyrimidine base pairs in a continuous stem capped by a structured pentaloop. The DKC mutant hairpin was observed to be slightly more stable than the wild-type hairpin, further shifting the pseudoknot-hairpin equilibrium to favor the mutant P2b hairpin. Here we examined the solution structure of the DKC mutant hairpin to identify the reason for this additional stability. We found that the mutant hairpin forms the same stem structure as wild-type and that the additional stabilization observed using optical melting can be explained by the formation of a YNMG-type tetraloop structure, with the last nucleotide of the pentaloop bulged out into the major groove. Our results provide a structural explanation for the increased stability of the mutant hairpin and further our understanding of the effect of this mutation on the structure and stability of the dominant conformation of the pseudoknot domain in this type of DKC.
Organizational Affiliation:
Department of Chemistry and Biochemistry, and Molecular Biology Institute, University of California, Los Angeles, Los Angeles, California 90095, USA.