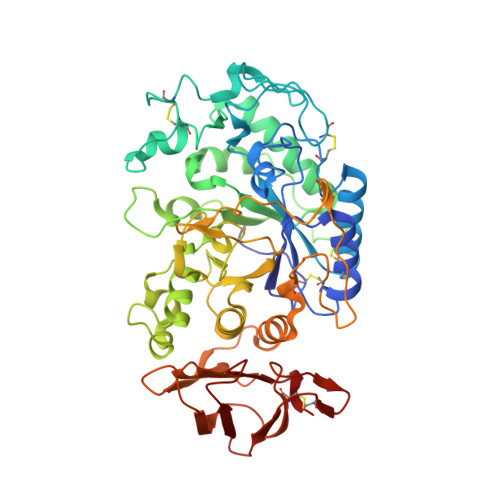
Structural studies of a Phe256Trp mutant of human salivary alpha-amylase: implications for the role of a conserved water molecule in enzyme activity
Ramasubbu, N., Sundar, K., Ragunath, C., Rafi, M.M.(2004) Arch Biochem Biophys 421: 115-124
- PubMed: 14678792
- DOI: https://doi.org/10.1016/j.abb.2003.10.007
- Primary Citation of Related Structures:
1Q4N - PubMed Abstract:
In the mechanism of hydrolysis of starch by alpha-amylases, a conserved water molecule bridging two catalytic residues has been implicated. In human salivary alpha-amylase (HSAmy), this water (W641), observed in many alpha-amylase structures, is part of a chain of water molecules. To test the hypothesis that W641 may be involved in the mechanism, Phe256 in the close vicinity was mutated to a Trp residue. X-ray structure of F256W complexed to 2-amino-2-(hydroxyethyl)-1,3-propanediol at 2.1A revealed that the water chain is disrupted. In the F256W structure exhibits a positional shift in His305, characteristic of alpha-amylase complex structures. Kinetic analysis, in comparison with HSAmy, revealed that the mutant exhibited a 70-fold decrease in the specific activity for starch and significantly reduced k(cat) (20-fold) and K(m) (4-fold) for maltoheptaoside. Collectively, these results suggest that W641 and the chain of water molecules may be critical for the alpha-amylase activity.
Organizational Affiliation:
Department of Oral Biology, New Jersey Dental School, University of Medicine and Dentistry, Newark, NJ 07103, USA. n.ramasubbu@umdnj.edu