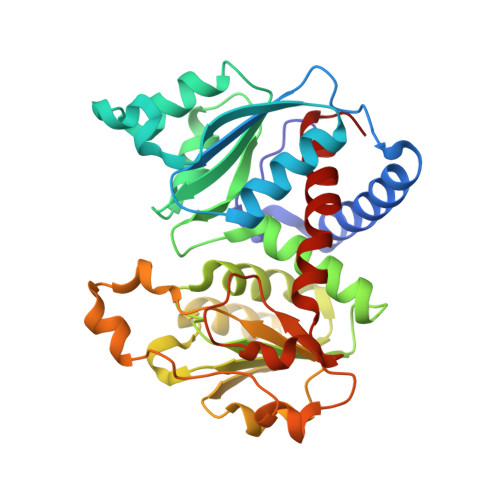
Aspartate Transcarbamylase from the Hyperthermophilic Archaeon Pyrococcus abyssi: Thermostability and 1.8A Resolution Crystal Structure of the Catalytic Subunit Complexed With the Bisubstrate Analogue N-Phosphonacetyl-L-aspartate.
Van Boxstael, S., Cunin, R., Khan, S., Maes, D.(2003) J Mol Biol 326: 203-216
- PubMed: 12547202
- DOI: https://doi.org/10.1016/s0022-2836(02)01228-7
- Primary Citation of Related Structures:
1ML4 - PubMed Abstract:
The Pyrococcus abyssi aspartate transcarbamylase (ATCase) shows a high degree of structural conservation with respect to the well-studied mesophilic Escherichia coli ATCase, including the association of catalytic and regulatory subunits. The adaptation of its catalytic function to high temperature was investigated, using enzyme purified from recombinant E.coli cells. At 90 degrees C, the activity of the trimeric catalytic subunit was shown to be intrinsically thermostable. Significant extrinsic stabilization by phosphate, a product of the reaction, was observed when the temperature was raised to 98 degrees C. Comparison with the holoenzyme showed that association with regulatory subunits further increases thermostability. To provide further insight into the mechanisms of its adaptation to high temperature, the crystal structure of the catalytic subunit liganded with the analogue N-phosphonacetyl-L-aspartate (PALA) was solved to 1.8A resolution and compared to that of the PALA-liganded catalytic subunit from E.coli. Interactions with PALA are strictly conserved. This, together with the similar activation energies calculated for the two proteins, suggests that the reaction mechanism of the P.abyssi catalytic subunit is similar to that of the E.coli subunit. Several structural elements potentially contributing to thermostability were identified: (i) a marked decrease in the number of thermolabile residues; (ii) an increased number of charged residues and a concomitant increase of salt links at the interface between the monomers, as well as the formation of an ion-pair network at the protein surface; (iii) the shortening of three loops and the shortening of the N and C termini. Other known thermostabilizing devices such as increased packing density or reduction of cavity volumes do not appear to contribute to the high thermostability of the P.abyssi enzyme.
Organizational Affiliation:
Laboratorium voor Erfelijkheidsleer en Microbiologie, Faculteit der Wetenschappen, Vrije Universiteit Brussel (VUB), 1 E. Gryson ave, B-1070, Brussels, Belgium.