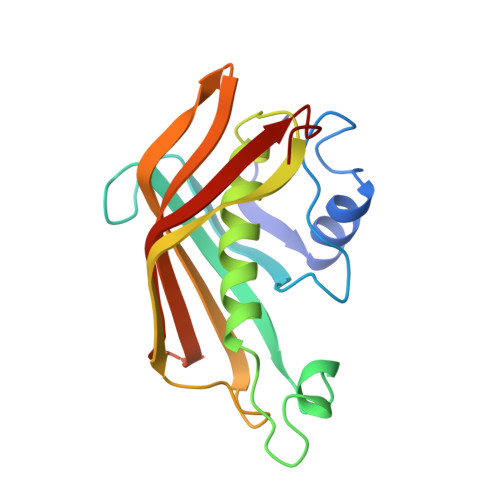
Structure of a dehydratase-isomerase from the bacterial pathway for biosynthesis of unsaturated fatty acids: two catalytic activities in one active site.
Leesong, M., Henderson, B.S., Gillig, J.R., Schwab, J.M., Smith, J.L.(1996) Structure 4: 253-264
- PubMed: 8805534
- DOI: https://doi.org/10.1016/s0969-2126(96)00030-5
- Primary Citation of Related Structures:
1MKA, 1MKB - PubMed Abstract:
Escherichia coli beta-hydroxydecanoyl thiol ester dehydrase (dehydrase) is essential to the biosynthesis of unsaturated fatty acids, by shunting a 10-carbon intermediate from the saturated fatty acid pathway into the unsaturated fatty acid pathway. Dehydrase catalyzes reactions of dehydration and of double-bond isomerization on 10-carbon thiol esters of acyl carrier protein (ACP). The aim of this work is to elucidate mechanisms for the two enzymatic reactions, which occur in an unusual bifunctional active site, and to understand the specificity of the enzyme for substrates with 10-carbon fatty acyl chains. Crystal structures at 2.0 A resolution for free dehydrase and for the enzyme modified by its classic, mechanism-based inactivator, 3-decynoyl-N-acetylcysteamine, have been determined. Dehydrase is a symmetric dimer with an unusual alpha+beta 'hot dog' fold. Each of the two independent active sites is located between the two subunits of the enzyme, and is a tunnel-shaped pocket completely isolated from the general solvent. Side chains of histidine from one subunit and aspartic acid from the other are the only potentially reactive protein groups in the active site. A two-base mechanism by which the histidine and aspartic acid together catalyze dehydration and isomerization reactions is consistent with the active-site structure. The unique topology of the protein fold and the identification of the active-site components reveal features of predictive value for another enzyme, FabZ, which may be the non-specific dehydratase involved in elongation of fatty acyl chains. A positively charged area surrounding the entrance to the active site, which could interact with the negatively charged ACP, was also found.
Organizational Affiliation:
Department of Biological Sciences, Purdue University, West Lafayette, IN 47907, USA.