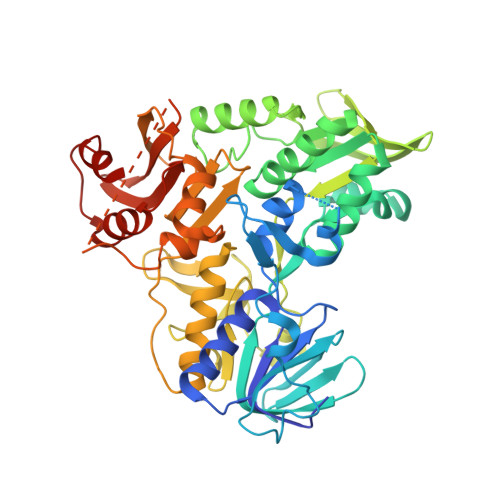
DNA binding is required for the apoptogenic action of apoptosis inducing factor.
Ye, H., Cande, C., Stephanou, N.C., Jiang, S., Gurbuxani, S., Larochette, N., Daugas, E., Garrido, C., Kroemer, G., Wu, H.(2002) Nat Struct Biol 9: 680-684
- PubMed: 12198487
- DOI: https://doi.org/10.1038/nsb836
- Primary Citation of Related Structures:
1M6I - PubMed Abstract:
The execution of apoptosis or programmed cell death comprises both caspase-dependent and caspase-independent processes. Apoptosis inducing factor (AIF) was identified as a major player in caspase-independent cell death. It induces chromatin condensation and initial DNA cleavage via an unknown molecular mechanism. Here we report the crystal structure of human AIF at 1.8 A resolution. The structure reveals the presence of a strong positive electrostatic potential at the AIF surface, although the calculated isoelectric point for the entire protein is neutral. We show that recombinant AIF interacts with DNA in a sequence-independent manner. In addition, in cells treated with an apoptotic stimulus, endogenous AIF becomes co-localized with DNA at an early stage of nuclear morphological changes. Structure-based mutagenesis shows that DNA-binding defective mutants of AIF fail to induce cell death while retaining nuclear translocation. The potential DNA-binding site identified from mutagenesis also coincides with computational docking of a DNA duplex. These observations suggest that AIF-induced nuclear apoptosis requires a direct interaction with DNA.
Organizational Affiliation:
Department of Biochemistry, Weill Medical College of Cornell University, 1300 York Avenue, New York, New York 10021, USA.