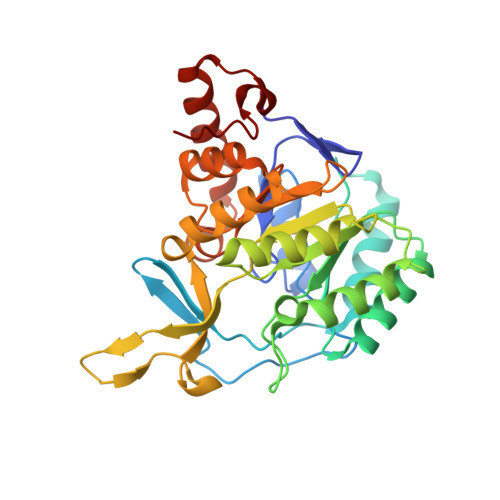
Lactococcus lactis dihydroorotate dehydrogenase A mutants reveal important facets of the enzymatic function
Norager, S., Arent, S., Bjornberg, O., Ottosen, M., Lo Leggio, L., Jensen, K.F., Larsen, S.(2003) J Biol Chem 278: 28812-28822
- PubMed: 12732650
- DOI: https://doi.org/10.1074/jbc.M303767200
- Primary Citation of Related Structures:
1JQV, 1JQX, 1JRB, 1JRC, 1JUB, 1JUE, 1OVD - PubMed Abstract:
Dihydroorotate dehydrogenases (DHODs) are flavoenzymes catalyzing the oxidation of (S)-dihydroorotate to orotate in the biosynthesis of UMP, the precursor of all other pyrimidine nucleotides. On the basis of sequence, DHODs can be divided into two classes, class 1, further divided in subclasses 1A and 1B, and class 2. This division corresponds to differences in cellular location and the nature of the electron acceptor. Herein we report a study of Lactococcus lactis DHODA, a representative of the class 1A enzymes. Based on the DHODA structure we selected seven residues that are highly conserved between both main classes of DHODs as well as three residues representing surface charges close to the active site for site-directed mutagenesis. The availability of both kinetic and structural data on the mutant enzymes allowed us to define the roles individual structural segments play in catalysis. We have also structurally proven the presence of an open active site loop in DHODA and obtained information about the interactions that control movements of loops around the active site. Furthermore, in one mutant structure we observed differences between the two monomers of the dimer, confirming an apparent asymmetry between the two substrate binding sites that was indicated by the kinetic results.
Organizational Affiliation:
Centre for Crystallographic Studies, University of Copenhagen, Universitetsparken 5, DK-2100 Copenhagen, Denmark.