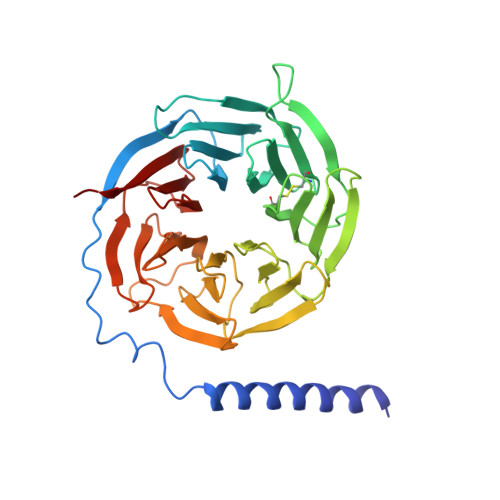
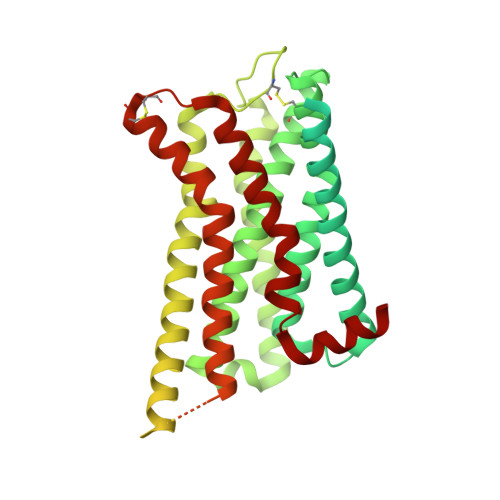
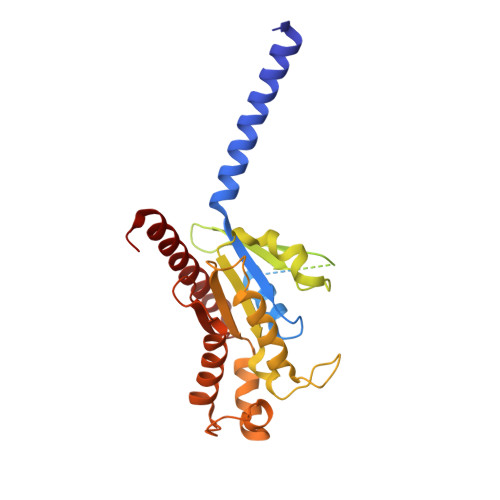
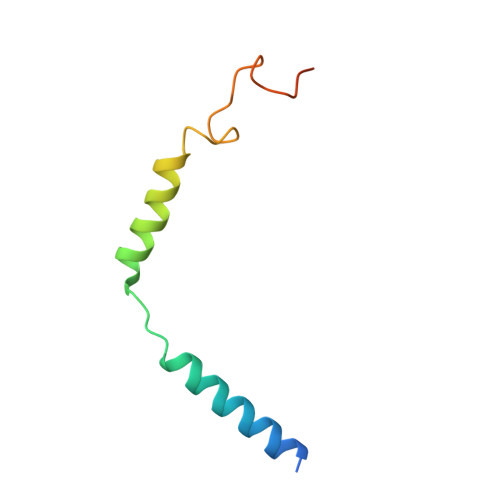
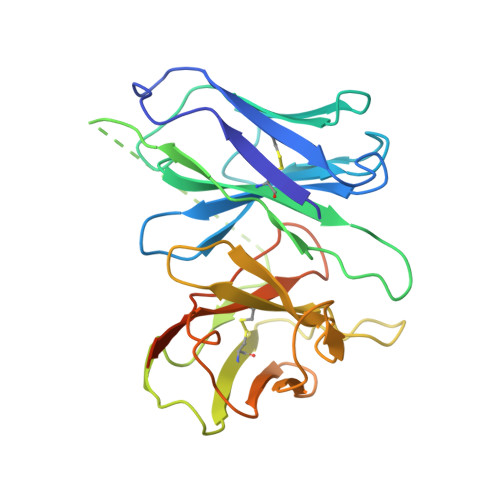
Structural insights into the human D1 and D2 dopamine receptor signaling complexes.
Zhuang, Y., Xu, P., Mao, C., Wang, L., Krumm, B., Zhou, X.E., Huang, S., Liu, H., Cheng, X., Huang, X.P., Shen, D.D., Xu, T., Liu, Y.F., Wang, Y., Guo, J., Jiang, Y., Jiang, H., Melcher, K., Roth, B.L., Zhang, Y., Zhang, C., Xu, H.E.(2021) Cell 184: 931-942.e18
- PubMed: 33571431
- DOI: https://doi.org/10.1016/j.cell.2021.01.027
- Primary Citation of Related Structures:
7JV5, 7JVP, 7JVQ, 7JVR - PubMed Abstract:
The D1- and D2-dopamine receptors (D1R and D2R), which signal through G s and G i , respectively, represent the principal stimulatory and inhibitory dopamine receptors in the central nervous system. D1R and D2R also represent the main therapeutic targets for Parkinson's disease, schizophrenia, and many other neuropsychiatric disorders, and insight into their signaling is essential for understanding both therapeutic and side effects of dopaminergic drugs. Here, we report four cryoelectron microscopy (cryo-EM) structures of D1R-G s and D2R-G i signaling complexes with selective and non-selective dopamine agonists, including two currently used anti-Parkinson's disease drugs, apomorphine and bromocriptine. These structures, together with mutagenesis studies, reveal the conserved binding mode of dopamine agonists, the unique pocket topology underlying ligand selectivity, the conformational changes in receptor activation, and potential structural determinants for G protein-coupling selectivity. These results provide both a molecular understanding of dopamine signaling and multiple structural templates for drug design targeting the dopaminergic system.
Organizational Affiliation:
The CAS Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai 201203, China; University of Chinese Academy of Sciences, Beijing 100049, China.