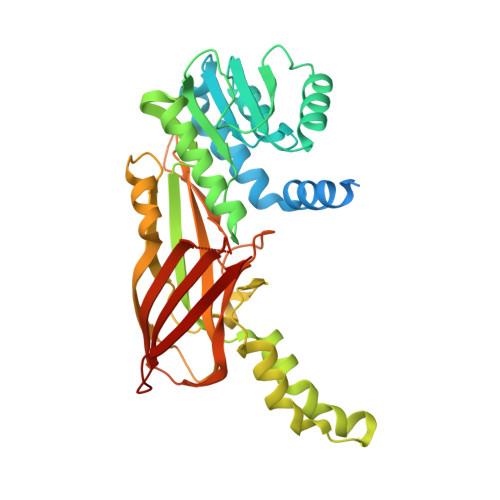
Crystal Structure of Human Protein arginine N-methyltransferase 6 (PRMT6) in complex with MT2739 inhibitor
Halabelian, L., Zeng, H., Dong, A., Schapira, M., De Freitas, R.F., Hutchinson, A., Seitova, A., Bountra, C., Edwards, A.M., Arrowsmith, C.H., Brown, P.J., Structural Genomics Consortium (SGC)To be published.