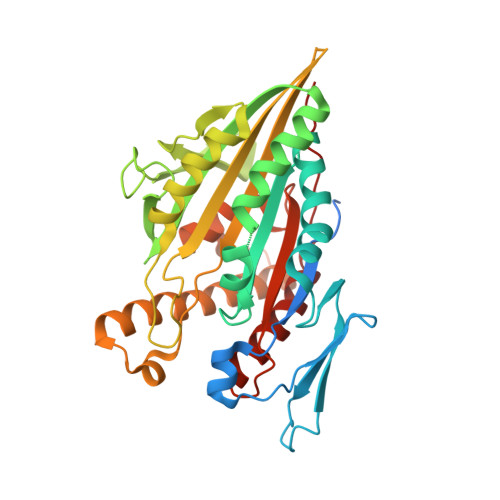
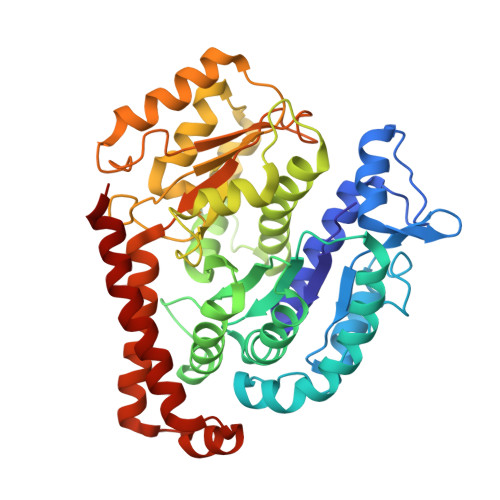
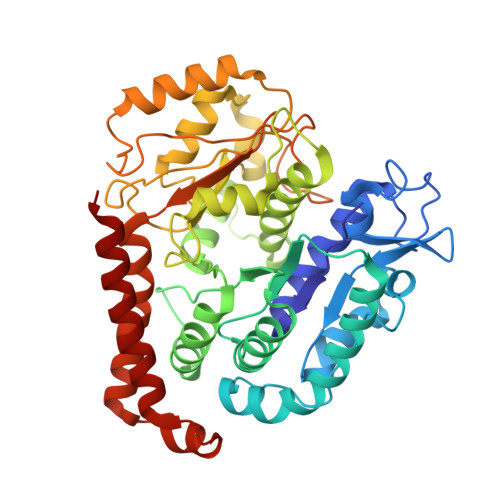
Structure of Microtubule-Trapped Human Kinesin-5 and Its Mechanism of Inhibition Revealed Using Cryoelectron Microscopy.
Pena, A., Sweeney, A., Cook, A.D., Topf, M., Moores, C.A.(2020) Structure 28: 450-457.e5
- PubMed: 32084356
- DOI: https://doi.org/10.1016/j.str.2020.01.013
- Primary Citation of Related Structures:
6TA3, 6TA4, 6TIW - PubMed Abstract:
Kinesin-5 motors are vital mitotic spindle components, and disruption of their function perturbs cell division. We investigated the molecular mechanism of the human kinesin-5 inhibitor GSK-1, which allosterically promotes tight microtubule binding. GSK-1 inhibits monomeric human kinesin-5 ATPase and microtubule gliding activities, and promotes the motor's microtubule stabilization activity. Using cryoelectron microscopy, we determined the 3D structure of the microtubule-bound motor-GSK-1 at 3.8 Å overall resolution. The structure reveals that GSK-1 stabilizes the microtubule binding surface of the motor in an ATP-like conformation, while destabilizing regions of the motor around the empty nucleotide binding pocket. Density corresponding to GSK-1 is located between helix-α4 and helix-α6 in the motor domain at its interface with the microtubule. Using a combination of difference mapping and protein-ligand docking, we characterized the kinesin-5-GSK-1 interaction and further validated this binding site using mutagenesis. This work opens up new avenues of investigation of kinesin inhibition and spindle perturbation.
Organizational Affiliation:
Institute of Structural and Molecular Biology, Birkbeck College, London WC1E 7HX, UK.