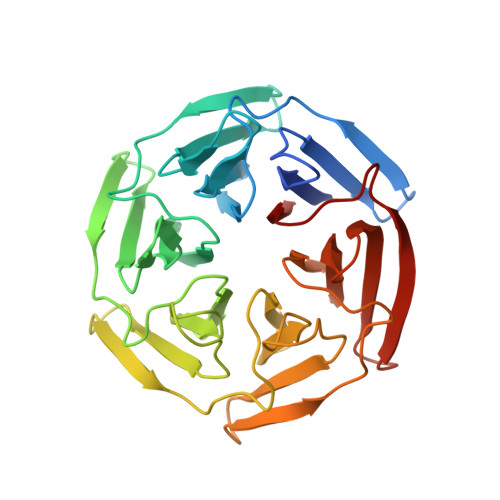
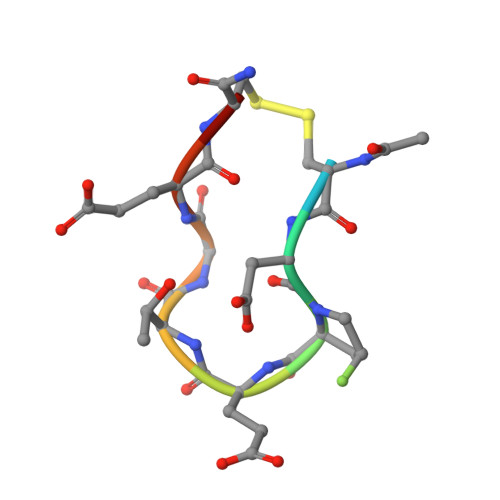
Optimization of linear and cyclic peptide inhibitors of KEAP1-NRF2 protein-protein interaction.
Colarusso, S., De Simone, D., Frattarelli, T., Andreini, M., Cerretani, M., Missineo, A., Moretti, D., Tambone, S., Kempf, G., Augustin, M., Steinbacher, S., Munoz-Sanjuan, I., Park, L., Summa, V., Tomei, L., Bresciani, A., Dominguez, C., Toledo-Sherman, L., Bianchi, E.(2020) Bioorg Med Chem 28: 115738-115738
- PubMed: 33065433
- DOI: https://doi.org/10.1016/j.bmc.2020.115738
- Primary Citation of Related Structures:
6T7V, 6T7Z - PubMed Abstract:
Inhibition of KEAP1-NRF2 protein-protein interaction is considered a promising strategy to selectively and effectively activate NRF2, a transcription factor which is involved in several pathologies such as Huntington's disease (HD). A library of linear peptides based on the NRF2-binding motifs was generated on the nonapeptide lead Ac-LDEETGEFL-NH 2 spanning residues 76-84 of the Neh2 domain of NRF2 with the aim to replace E78, E79 and E82 with non-acidic amino acids. A deeper understanding of the features and accessibility of the T80 subpocket was also targeted by structure-based design. Approaches to improve cell permeability were investigated using both different classes of cyclic peptides and conjugation to cell-penetrating peptides. This insight will guide future design of macrocycles, peptido-mimetics and, most importantly, small neutral brain-penetrating molecules to evaluate whether NRF2 activators have utility in HD.
Organizational Affiliation:
Department of Drug Discovery, IRBM Spa, Via Pontina km 30.600, 00071 Pomezia, Rome, Italy. Electronic address: s.colarusso@irbm.com.