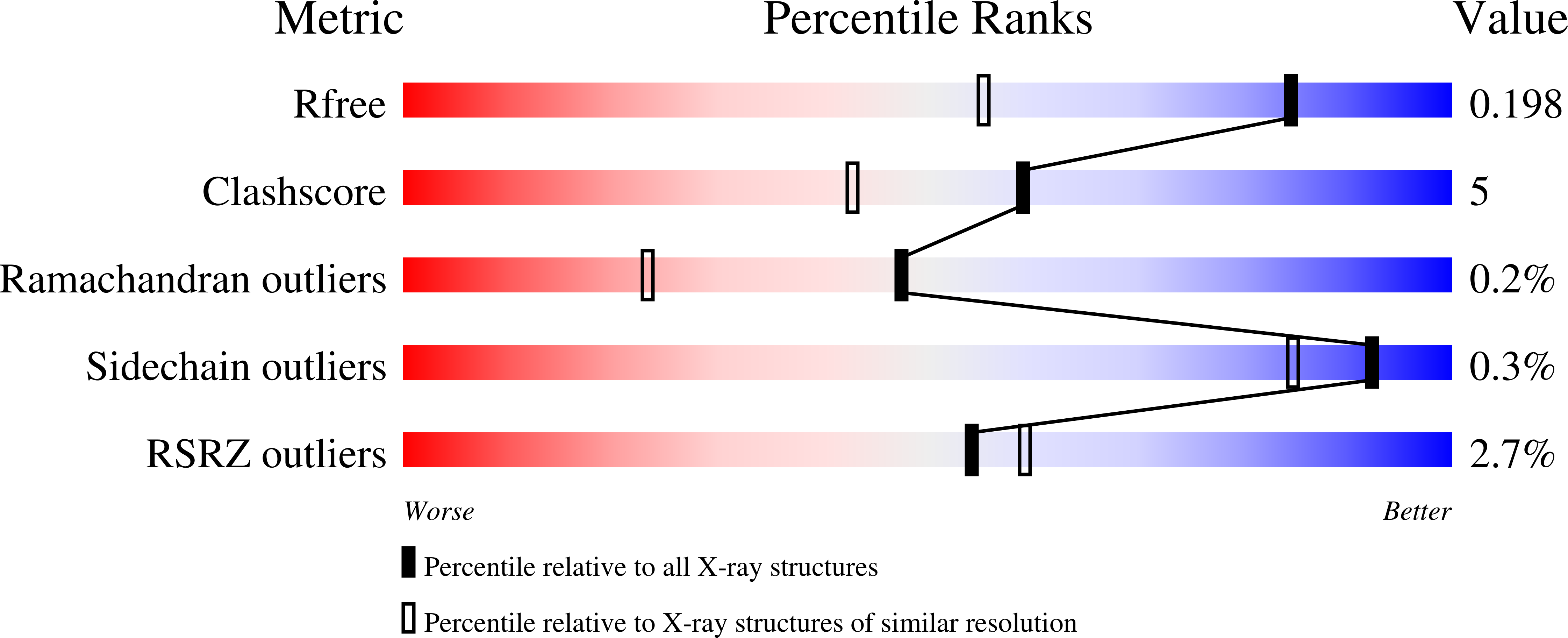
Covalent and noncovalent constraints yield a figure eight-like conformation of a peptide inhibiting the menin-MLL interaction.
Fortuna, P., Linhares, B.M., Purohit, T., Pollock, J., Cierpicki, T., Grembecka, J., Berlicki, L.(2020) Eur J Med Chem 207: 112748-112748
- PubMed: 32882610
- DOI: https://doi.org/10.1016/j.ejmech.2020.112748
- Primary Citation of Related Structures:
6OPJ - PubMed Abstract:
The interaction between menin and mixed lineage leukemia (MLL) was identified as an interesting target for treating some cancers including acute leukemia. On the basis of the known crystal structure of the MBM1-menin complex (MBM - menin binding motif), several cyclic peptides were designed. Elaboration of the effective cyclization strategy using a metathesis reaction allowed for a successfully large number of derivatives to be obtained. Subsequent optimization of the loop size, as well as N-terminal, central and C-terminal parts of the studied peptides resulted in structures exhibiting low nanomolar activities. A crystal structure of an inhibitor-menin complex revealed a compact conformation of the ligand molecule, which is stabilized not only by the introduction of a covalent linker but also three intramolecular hydrogen bonds. The inhibitor adopts a figure eight-like conformation, which perfectly fits the cleft of menin. We demonstrated that the development of compact, miniprotein-like structures is a highly effective approach for inhibition of protein-protein interactions.
Organizational Affiliation:
Department of Bioorganic Chemistry, Faculty of Chemistry, Wrocław University of Science and Technology, Wybrzeże Wyspiańskiego 27, 50-370, Wrocław, Poland.