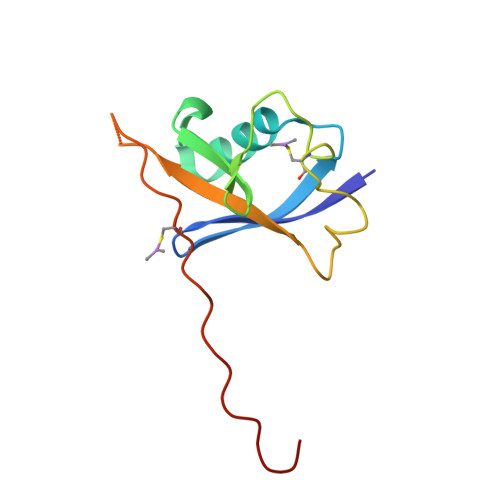
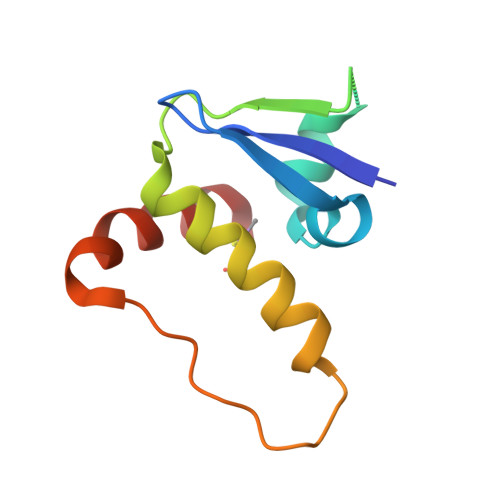
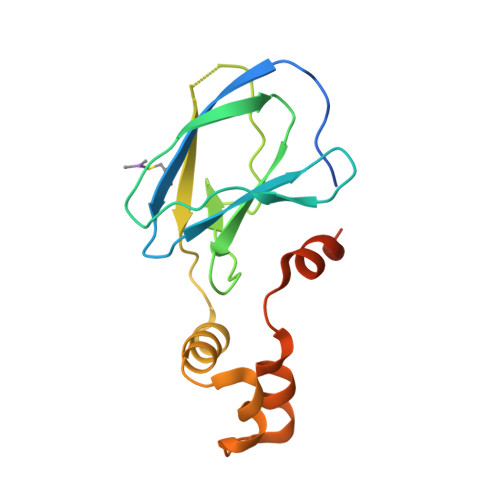
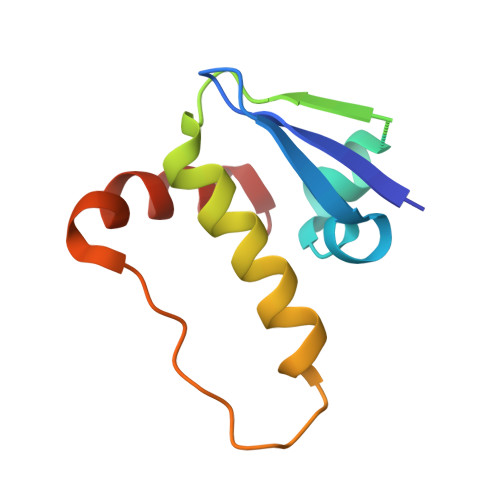
Surface Probing by Fragment-Based Screening and Computational Methods Identifies Ligandable Pockets on the von Hippel-Lindau (VHL) E3 Ubiquitin Ligase.
Lucas, X., Van Molle, I., Ciulli, A.(2018) J Med Chem 61: 7387-7393
- PubMed: 30040896
- DOI: https://doi.org/10.1021/acs.jmedchem.8b00842
- Primary Citation of Related Structures:
6GMN, 6GMQ, 6GMR, 6GMX - PubMed Abstract:
Beyond the targeting of E3 ubiquitin ligases to inhibit protein homeostasis, E3 ligase binders can be repurposed as targeted protein degraders (PROTACs or molecular glues). We sought to identify new binders of the VHL E3 ligase by biophysical fragment-based screening followed by X-ray crystallographic soaking. We identified fragments binding at the ElonginC:Cullin2 interface and a new cryptic pocket in VHL, along with other potential ligandable sites predicted computationally and found to bind solvent molecules in crystal structures. The elucidated interactions provide starting points for future ligand development.
Organizational Affiliation:
Division of Biological Chemistry and Drug Discovery, James Black Centre, School of Life Sciences , University of Dundee , Dow Street , Dundee , DD1 5EH , United Kingdom.