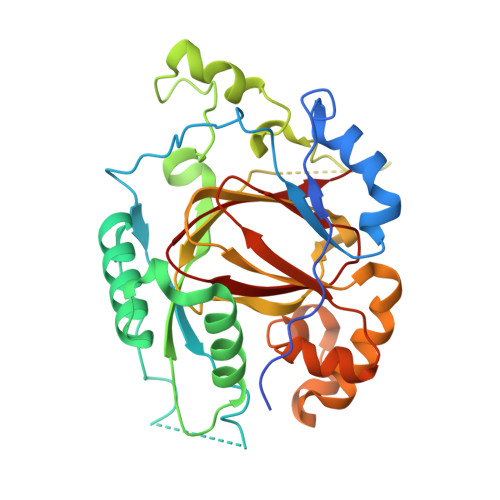
Structure-Based Engineering of Irreversible Inhibitors against Histone Lysine Demethylase KDM5A.
Horton, J.R., Woodcock, C.B., Chen, Q., Liu, X., Zhang, X., Shanks, J., Rai, G., Mott, B.T., Jansen, D.J., Kales, S.C., Henderson, M.J., Cyr, M., Pohida, K., Hu, X., Shah, P., Xu, X., Jadhav, A., Maloney, D.J., Hall, M.D., Simeonov, A., Fu, H., Vertino, P.M., Cheng, X.(2018) J Med Chem 61: 10588-10601
- PubMed: 30392349
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01219
- Primary Citation of Related Structures:
6DQ4, 6DQ5, 6DQ6, 6DQ8, 6DQ9, 6DQA, 6DQB - PubMed Abstract:
The active sites of hundreds of human α-ketoglutarate (αKG) and Fe(II)-dependent dioxygenases are exceedingly well preserved, which challenges the design of selective inhibitors. We identified a noncatalytic cysteine (Cys481 in KDM5A) near the active sites of KDM5 histone H3 lysine 4 demethylases, which is absent in other histone demethylase families, that could be explored for interaction with the cysteine-reactive electrophile acrylamide. We synthesized analogs of a thienopyridine-based inhibitor chemotype, namely, 2-((3-aminophenyl)(2-(piperidin-1-yl)ethoxy)methyl)thieno[3,2- b]pyridine-7-carboxylic acid (N70) and a derivative containing a (dimethylamino)but-2-enamido)phenyl moiety (N71) designed to form a covalent interaction with Cys481. We characterized the inhibitory and binding activities against KDM5A and determined the cocrystal structures of the catalytic domain of KDM5A in complex with N70 and N71. Whereas the noncovalent inhibitor N70 displayed αKG-competitive inhibition that could be reversed after dialysis, inhibition by N71 was dependent on enzyme concentration and persisted even after dialysis, consistent with covalent modification.
Organizational Affiliation:
Department of Molecular and Cellular Oncology , The University of Texas MD Anderson Cancer Center , Houston , Texas 77030 , United States.