Promutagenicity of 8-Chloroguanine, A Major Inflammation-Induced Halogenated DNA Lesion.
Kou, Y., Koag, M.C., Lee, S.(2019) Molecules 24
- PubMed: 31569643
- DOI: https://doi.org/10.3390/molecules24193507
- Primary Citation of Related Structures:
6CRH - PubMed Abstract:
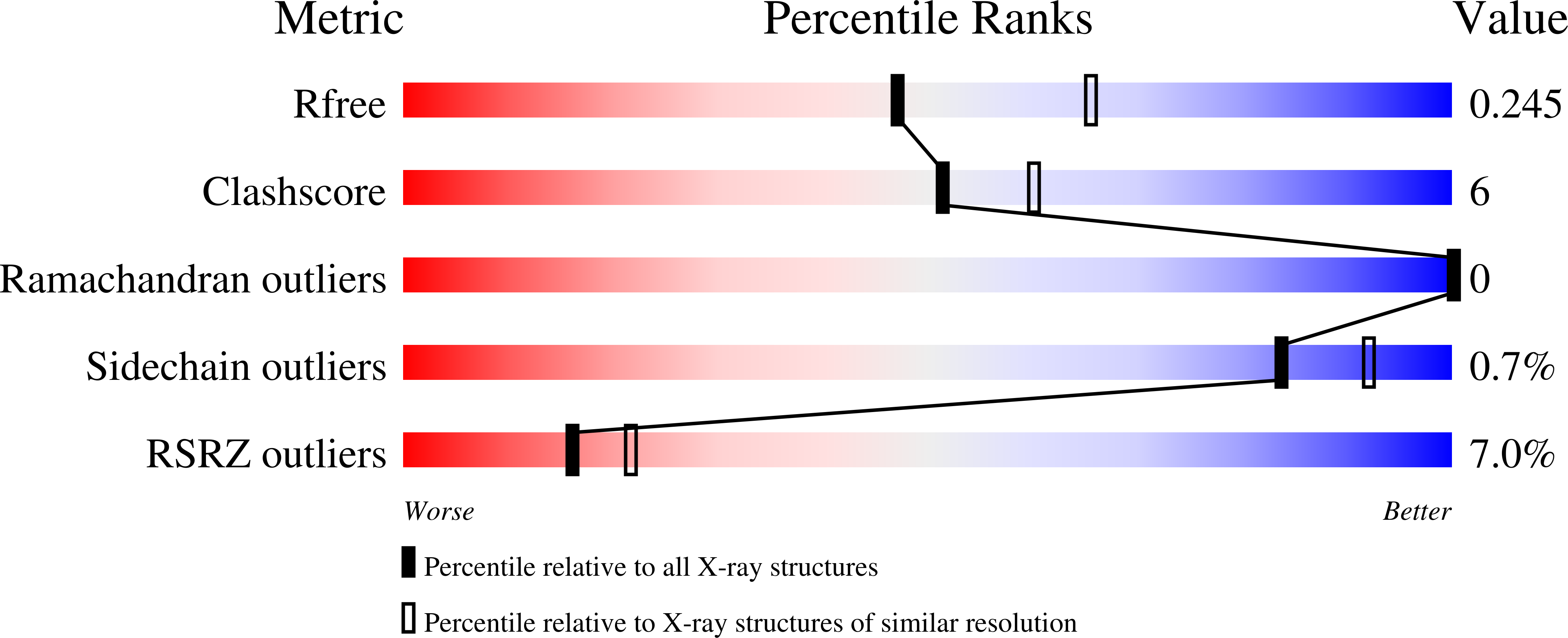
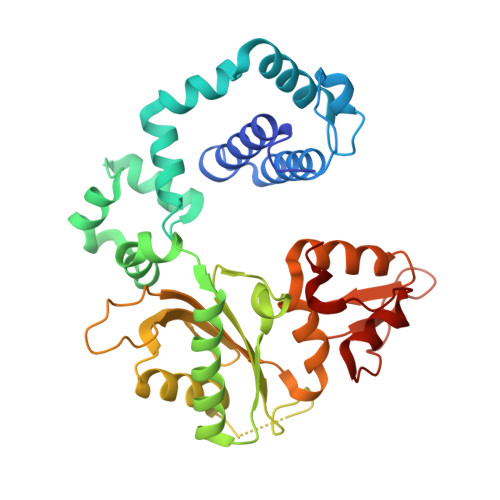
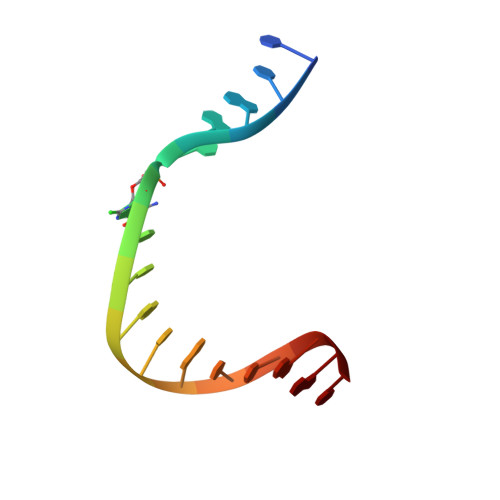
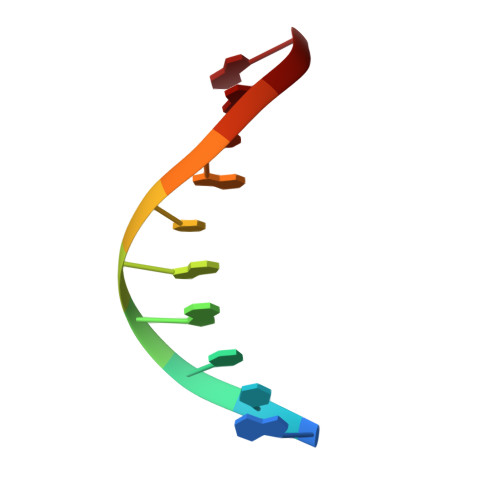
Chronic inflammation is closely associated with cancer development. One possible mechanism for inflammation-induced carcinogenesis is DNA damage caused by reactive halogen species, such as hypochlorous acid, which is released by myeloperoxidase to kill pathogens. Hypochlorous acid can attack genomic DNA to produce 8-chloro-2'-deoxyguanosine (ClG) as a major lesion. It has been postulated that ClG promotes mutagenic replication using its syn conformer; yet, the structural basis for ClG-induced mutagenesis is unknown. We obtained crystal structures and kinetics data for nucleotide incorporation past a templating ClG using human DNA polymerase β (polβ) as a model enzyme for high-fidelity DNA polymerases. The structures showed that ClG formed base pairs with incoming dCTP and dGTP using its anti and syn conformers, respectively. Kinetic studies showed that polβ incorporated dGTP only 15-fold less efficiently than dCTP, suggesting that replication across ClG is promutagenic. Two hydrogen bonds between syn -ClG and anti -dGTP and a water-mediated hydrogen bond appeared to facilitate mutagenic replication opposite the major halogenated guanine lesion. These results suggest that ClG in DNA promotes G to C transversion mutations by forming Hoogsteen base pairing between syn -ClG and anti -G during DNA synthesis.
Organizational Affiliation:
The Division of Chemical Biology and Medicinal Chemistry, College of Pharmacy, The University of Texas at Austin, 2409 University Avenue, Austin, TX 78712, USA. yik109@hotmail.com.