Crystal structures of monometallic dihydropyrimidinase and the human dihydroorotase domain K1556A mutant reveal no lysine carbamylation within the active site
Cheng, J.H., Huang, Y.H., Lin, J.J., Huang, C.Y.(2018) Biochem Biophys Res Commun 505: 439-444
- PubMed: 30268498
- DOI: https://doi.org/10.1016/j.bbrc.2018.09.153
- Primary Citation of Related Structures:
5YNZ, 6AJD - PubMed Abstract:
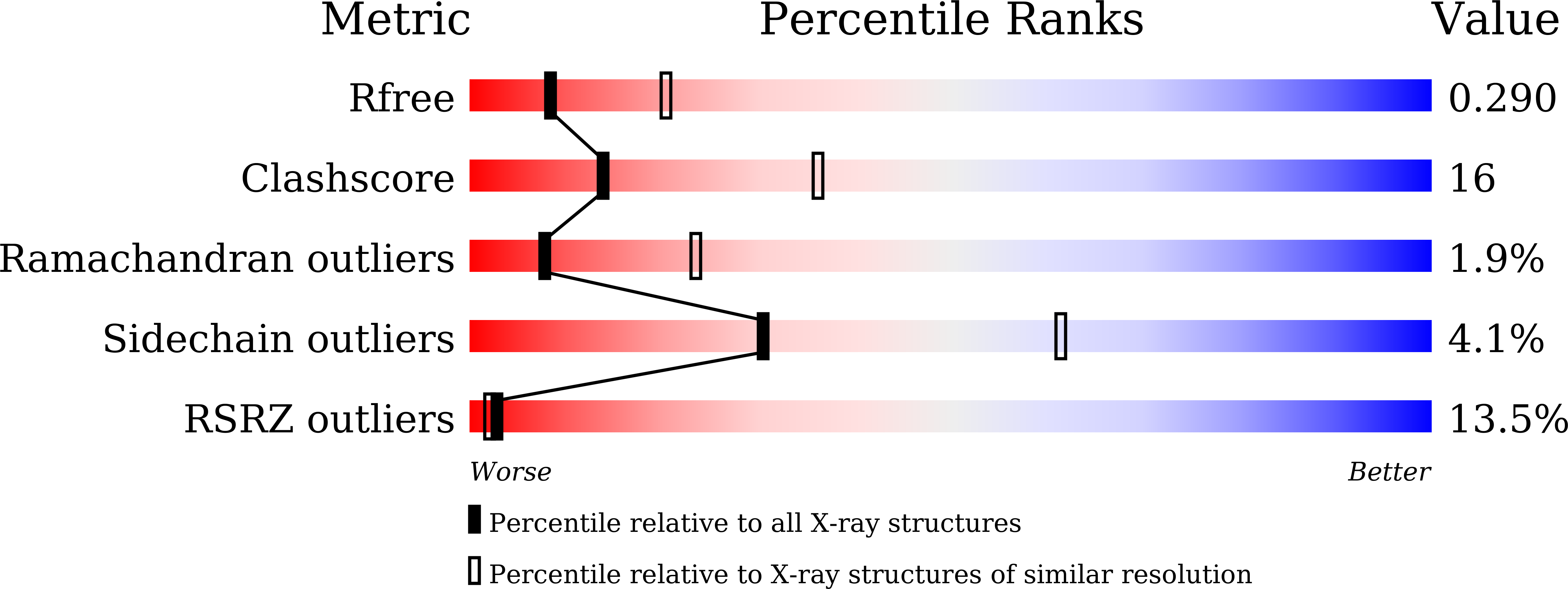
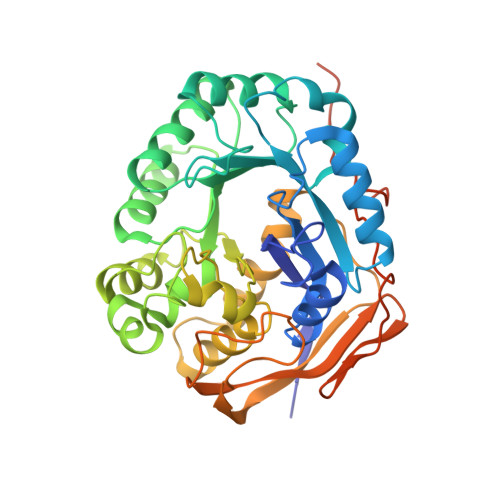
Dihydropyrimidinase (DHPase) is a member of the cyclic amidohydrolase family, which also includes allantoinase, dihydroorotase (DHOase), hydantoinase, and imidase. Almost all of these zinc metalloenzymes possess a binuclear metal center in which two metal ions are bridged by a post-translational carbamylated Lys. Crystal structure of Tetraodon nigroviridis DHPase reveals that one zinc ion is sufficient to stabilize Lys carbamylation. In this study, we found that one metal coordination was not sufficient to fix CO 2 to the Lys in bacterial DHPase. We prepared and characterized mono-Zn DHPase from Pseudomonas aeruginosa (PaDHPase), and the catalytic activity of mono-Zn PaDHPase was not detected. The crystal structure of mono-Zn PaDHPase determined at 2.23 Å resolution (PDB entry 6AJD) revealed that Lys150 was no longer carbamylated. This finding indicated the decarbamylation of the Lys during the metal chelating process. To confirm the state of Lys carbamylation in mono-Zn PaDHPase in solution, mass spectrometric (MS) analysis was carried out. The MS result was in agreement with the theoretical value for uncarbamylated PaDHPase. Crystal structure of the human DHOase domain (huDHOase) K1556A mutant was also determined (PDB entry 5YNZ), and the structure revealed that the active site of huDHOase K1556A mutant contained one metal ion. Like mono-Zn PaDHPase, oxygen ligands of the carbamylated Lys were not required for Znα binding. Considering the collective data from X-ray crystal structure and MS analysis, mono-Zn PaDHPase in both crystalline state and solution was not carbamylated. In addition, structural evidences indicated that post-translational carbamylated Lys was not required for Znα binding in PaDHPase and in huDHOase.
Organizational Affiliation:
School of Biomedical Sciences, Chung Shan Medical University, No.110, Sec.1, Chien-Kuo N. Rd., Taichung City, Taiwan.