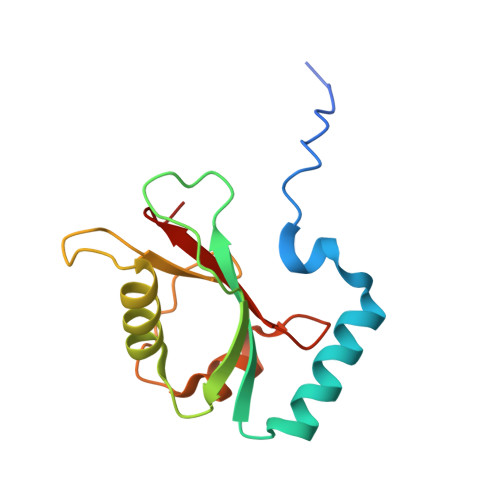
Insights into links between autophagy and the ubiquitin system from the structure of LC3B bound to the LIR motif from the E3 ligase NEDD4.
Qiu, Y., Zheng, Y., Wu, K.P., Schulman, B.A.(2017) Protein Sci 26: 1674-1680
- PubMed: 28470758
- DOI: https://doi.org/10.1002/pro.3186
- Primary Citation of Related Structures:
5V4K - PubMed Abstract:
Members of the LC3/GABARAP family of ubiquitin-like proteins function during autophagy by serving as membrane linked protein-binding platforms. Their C-termini are physically attached to membranes through covalent linkage to primary amines on lipids such as phosphatidylethanolamine, while their ubiquitin-like fold domains bind "LIR" (LC3-Interacting Region) sequences found within an extraordinarily diverse array of proteins including regulators of autophagy, adaptors that recruit ubiquitinated cargoes to be degraded, and even proteins controlling processes at membranes that are not associated with autophagy. Recently, LC3/GABARAP proteins were found to bind the ubiquitin E3 ligase NEDD4 to influence ubiquitination associated with autophagy in human cell lines. Here, we use purified recombinant proteins to define LC3B interactions with a specific LIR sequence from NEDD4, present a crystal structure showing atomic details of the interaction, and show that LC3B-binding can steer intrinsic NEDD4 E3 ligase activity. The data provide detailed molecular insights underlying recruitment of an E3 ubiquitin ligase to phagophores during autophagy.
Organizational Affiliation:
Department of Structural Biology, St. Jude Children's Research Hospital, Memphis, Tennessee, 38105.