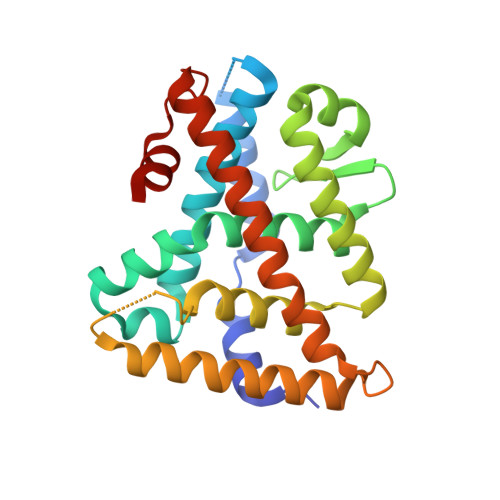
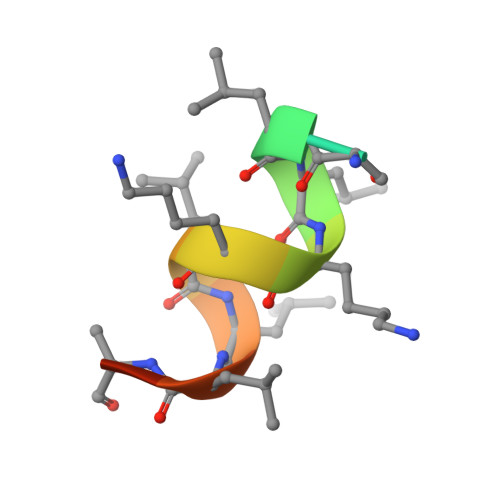
Structure and Dynamics of the Liver Receptor Homolog 1-PGC1 alpha Complex.
Mays, S.G., Okafor, C.D., Tuntland, M.L., Whitby, R.J., Dharmarajan, V., Stec, J., Griffin, P.R., Ortlund, E.A.(2017) Mol Pharmacol 92: 1-11
- PubMed: 28363985
- DOI: https://doi.org/10.1124/mol.117.108514
- Primary Citation of Related Structures:
5UNJ - PubMed Abstract:
Peroxisome proliferator-activated gamma coactivator 1- α (PGC1 α ) regulates energy metabolism by directly interacting with transcription factors to modulate gene expression. Among the PGC1 α binding partners is liver receptor homolog 1 (LRH-1; NR5A2), an orphan nuclear hormone receptor that controls lipid and glucose homeostasis. Although PGC1 α is known to bind and activate LRH-1, mechanisms through which PGC1 α changes LRH-1 conformation to drive transcription are unknown. Here, we used biochemical and structural methods to interrogate the LRH-1-PGC1 α complex. Purified, full-length LRH-1, as well as isolated ligand binding domain, bound to PGC1 α with higher affinity than to the coactivator, nuclear receptor coactivator-2 (Tif2), in coregulator peptide recruitment assays. We present the first crystal structure of the LRH-1-PGC1 α complex, which depicts several hydrophobic contacts and a strong charge clamp at the interface between these partners. In molecular dynamics simulations, PGC1 α induced correlated atomic motion throughout the entire LRH-1 activation function surface, which was dependent on charge-clamp formation. In contrast, Tif2 induced weaker signaling at the activation function surface than PGC1 α but promoted allosteric signaling from the helix 6/ β -sheet region of LRH-1 to the activation function surface. These studies are the first to probe mechanisms underlying the LRH-1-PGC1 α interaction and may illuminate strategies for selective therapeutic targeting of PGC1 α -dependent LRH-1 signaling pathways.
Organizational Affiliation:
Department of Biochemistry, Emory University School of Medicine, Atlanta, Georgia (S.G.M., C.D.O., M.L.T., E.A.O.); School of Chemistry, University of Southampton, Southampton, United Kingdom (R.J.W., J.S.); and Department of Molecular Medicine, Scripps Research Institute, Jupiter, Florida (V.D., P.R.G.).