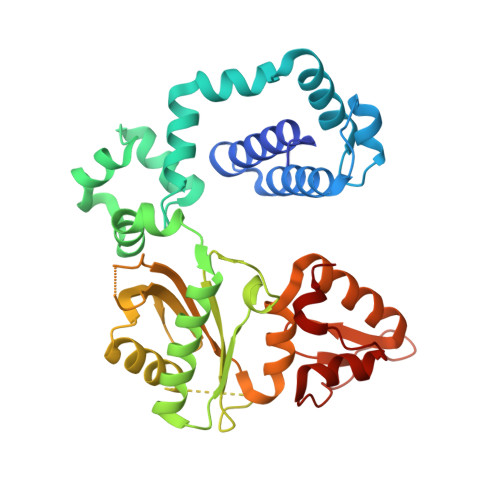
Remote Mutations Induce Functional Changes in Active Site Residues of Human DNA Polymerase beta.
Eckenroth, B.E., Towle-Weicksel, J.B., Nemec, A.A., Murphy, D.L., Sweasy, J.B., Doublie, S.(2017) Biochemistry 56: 2363-2371
- PubMed: 28402631
- DOI: https://doi.org/10.1021/acs.biochem.6b01287
- Primary Citation of Related Structures:
5U8G, 5U8H, 5U8I - PubMed Abstract:
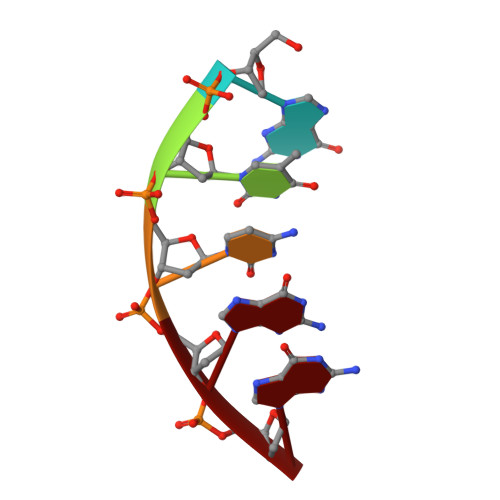
With the formidable growth in the volume of genetic information, it has become essential to identify and characterize mutations in macromolecules not only to predict contributions to disease processes but also to guide the design of therapeutic strategies. While mutations of certain residues have a predictable phenotype based on their chemical nature and known structural position, many types of mutations evade prediction based on current information. Described in this work are the crystal structures of two cancer variants located in the palm domain of DNA polymerase β (pol β), S229L and G231D, whose biological phenotype was not readily linked to a predictable structural implication. Structural results demonstrate that the mutations elicit their effect through subtle influences on secondary interactions with a residue neighboring the active site. Residues 229 and 231 are 7.5 and 12.5 Å, respectively, from the nearest active site residue, with a β-strand between them. A residue on this intervening strand, M236, appears to transmit fine structural perturbations to the catalytic metal-coordinating residue D256, affecting its conformational stability.
Organizational Affiliation:
Department of Microbiology and Molecular Genetics, University of Vermont , Stafford Hall, 95 Carrigan Drive, Burlington, Vermont 05405, United States.