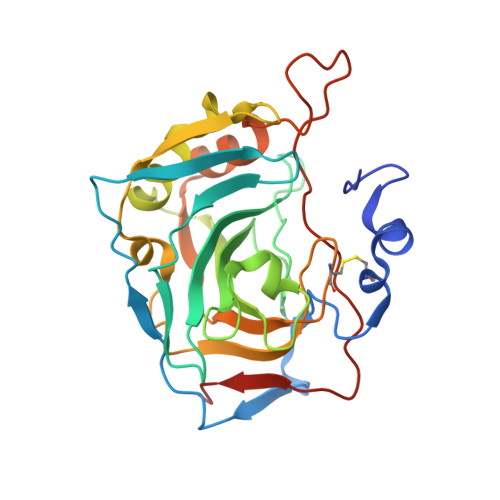
Combinatorial Design of Isoform-Selective N-Alkylated Benzimidazole-Based Inhibitors of Carbonic Anhydrases
Capkauskaite, E., Linkuviene, V., Smirnov, A., Milinaviciute, G., Timm, D., Kasiliauskaite, A., Manakova, E., Grazulis, S., Matulis, D.(2017) ChemistrySelect