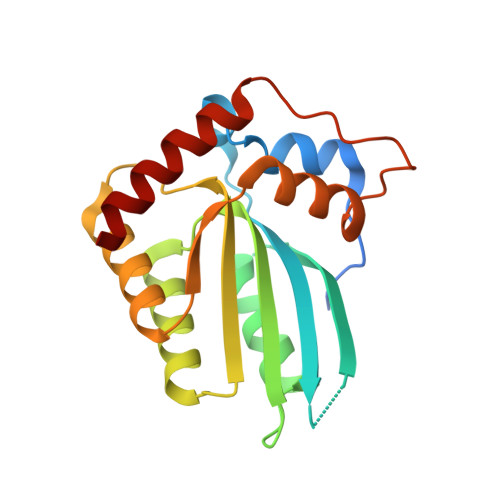
1.92 Angstrom Zinc-Free APOBEC3F Catalytic Domain Crystal Structure.
Shaban, N.M., Shi, K., Li, M., Aihara, H., Harris, R.S.(2016) J Mol Biol 428: 2307-2316
- PubMed: 27139641
- DOI: https://doi.org/10.1016/j.jmb.2016.04.026
- Primary Citation of Related Structures:
5HX4, 5HX5 - PubMed Abstract:
The APOBEC3 family of DNA cytosine deaminases is capable of restricting the replication of HIV-1 and other pathogens. Here, we report a 1.92 Å resolution crystal structure of the Vif-binding and catalytic domain of APOBEC3F (A3F). This structure is distinct from the previously published APOBEC and phylogenetically related deaminase structures, as it is the first without zinc in the active site. We determined an additional structure containing zinc in the same crystal form that allows direct comparison with the zinc-free structure. In the absence of zinc, the conserved active site residues that normally participate in zinc coordination show unique conformations, including a 90 degree rotation of His249 and disulfide bond formation between Cys280 and Cys283. We found that zinc coordination is influenced by pH, and treating the protein at low pH in crystallization buffer is sufficient to remove zinc. Zinc coordination and catalytic activity are reconstituted with the addition of zinc only in a reduced environment likely due to the two active site cysteines readily forming a disulfide bond when not coordinating zinc. We show that the enzyme is active in the presence of zinc and cobalt but not with other divalent metals. These results unexpectedly demonstrate that zinc is not required for the structural integrity of A3F and suggest that metal coordination may be a strategy for regulating the activity of A3F and related deaminases.
Organizational Affiliation:
Department of Biochemistry, Molecular Biology, and Biophysics, Masonic Cancer Center, Institute for Molecular Virology, University of Minnesota, Minneapolis, MN 55455, USA. Electronic address: nmshaban@umn.edu.