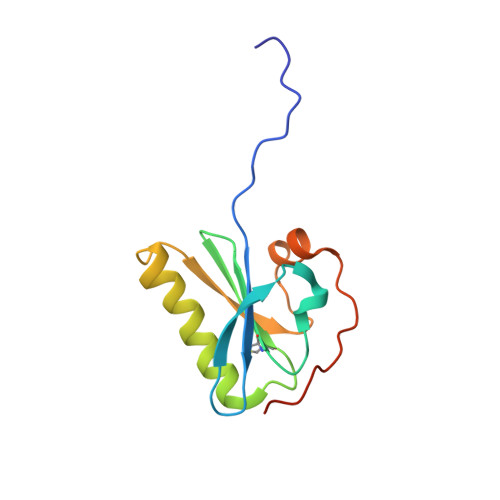
Molecular basis of a novel renal amyloidosis due to N184K gelsolin variant.
Boni, F., Milani, M., Porcari, R., Barbiroli, A., Ricagno, S., de Rosa, M.(2016) Sci Rep 6: 33463-33463
- PubMed: 27633054
- DOI: https://doi.org/10.1038/srep33463
- Primary Citation of Related Structures:
5FAE, 5FAF - PubMed Abstract:
Mutations in gelsolin are responsible for a systemic amyloidosis first described in 1969. Until recently, the disease was associated with two substitutions of the same residue, leading to the loss of the calcium binding site. Novel interest arose in 2014 when the N184K variant of the protein was identified as the etiological agent of a novel kidney-localized amyloidosis. Here we provide a first rationale for N184K pathogenicity. We show that the mutation induces a destabilization of gelsolin second domain, without compromising its calcium binding capacity. X-ray data combined with molecular dynamics simulations demonstrates that the primary source of the destabilization is a loss of connectivity in proximity of the metal. Such rearrangement of the H-bond network does not have a major impact on the overall fold of the domain, nevertheless, it increases the flexibility of a stretch of the protein, which is consequently processed by furin protease. Overall our data suggest that the N184K variant is subjected to the same aberrant proteolytic events responsible for the formation of amyloidogenic fragments in the previously characterized mutants. At the same time our data suggest that a broader number of mutations, unrelated to the metal binding site, can lead to a pathogenic phenotype.
Organizational Affiliation:
CNR Istituto di Biofisica, c/o Dipartimento di Bioscienze, Università degli Studi di Milano, 20133 Milano, Italy.