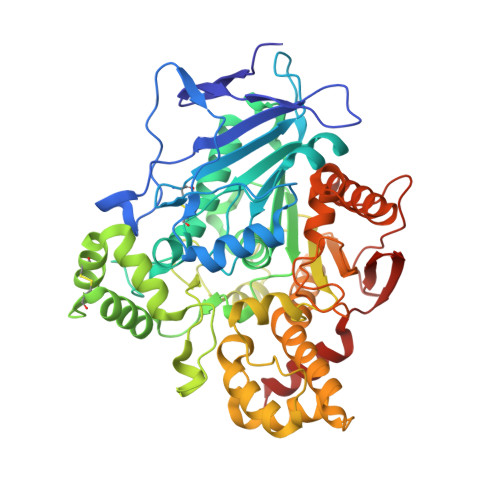
Comparison of the Structure and Activity of Glycosylated and Aglycosylated Human Carboxylesterase 1.
Arena De Souza, V., Scott, D.J., Nettleship, J.E., Rahman, N., Charlton, M.H., Walsh, M.A., Owens, R.J.(2015) PLoS One 10: 43919
- PubMed: 26657071
- DOI: https://doi.org/10.1371/journal.pone.0143919
- Primary Citation of Related Structures:
5A7F, 5A7G, 5A7H - PubMed Abstract:
Human Carboxylesterase 1 (hCES1) is the key liver microsomal enzyme responsible for detoxification and metabolism of a variety of clinical drugs. To analyse the role of the single N-linked glycan on the structure and activity of the enzyme, authentically glycosylated and aglycosylated hCES1, generated by mutating asparagine 79 to glutamine, were produced in human embryonic kidney cells. Purified enzymes were shown to be predominantly trimeric in solution by analytical ultracentrifugation. The purified aglycosylated enzyme was found to be more active than glycosylated hCES1 and analysis of enzyme kinetics revealed that both enzymes exhibit positive cooperativity. Crystal structures of hCES1 a catalytically inactive mutant (S221A) and the aglycosylated enzyme were determined in the absence of any ligand or substrate to high resolutions (1.86 Å, 1.48 Å and 2.01 Å, respectively). Superposition of all three structures showed only minor conformational differences with a root mean square deviations of around 0.5 Å over all Cα positions. Comparison of the active sites of these un-liganded enzymes with the structures of hCES1-ligand complexes showed that side-chains of the catalytic triad were pre-disposed for substrate binding. Overall the results indicate that preventing N-glycosylation of hCES1 does not significantly affect the structure or activity of the enzyme.
Organizational Affiliation:
UK OPPF-UK, The Research Complex at Harwell, Rutherford Appleton Laboratory Harwell Oxford, Oxfordshire, United Kingdom.