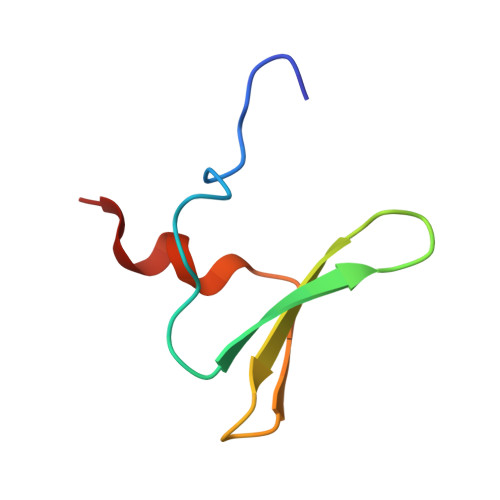
Crystal structure of the first WW domain of human YAP2 isoform.
Martinez-Rodriguez, S., Bacarizo, J., Luque, I., Camara-Artigas, A.(2015) J Struct Biol 191: 381-387
- PubMed: 26256245
- DOI: https://doi.org/10.1016/j.jsb.2015.08.001
- Primary Citation of Related Structures:
4REX - PubMed Abstract:
The WW domains are the smallest modular domains known. The study of the structural basis of their stability is important to understand their physiological role. These domains are intrinsically flexible, which makes them difficult to crystallize. The first WW domain of the human Yes tyrosine kinase Associated Protein (YAP) has been crystallized and its structure has been solved by X-ray diffraction at 1.6 Å resolution. Crystals belong to the orthorhombic space group P21212 with unit cell parameters a=42.67, b=43.10 and c=21.30. The addition of proline and other small-molecule additives improves drastically the quality of the crystals. The interactions that stabilize this minimal modular domain have been analysed. This crystal structure reveals that, besides the stabilization of the hydrophobic core of the protein by the aromatic cluster formed by Trp177-Phe189-Pro202, some salt-bridges interactions might affect the stability of the domain.
Organizational Affiliation:
Department of Physical Chemistry, University of Granada, Avda. de Fuentenueva s/n, Granada 18071, Spain.