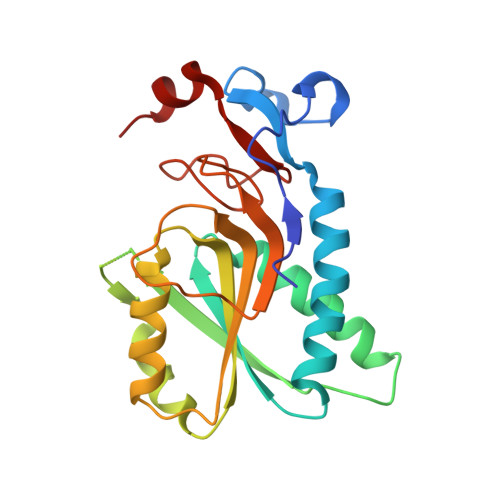
Aza-acyclic Nucleoside Phosphonates Containing a Second Phosphonate Group As Inhibitors of the Human, Plasmodium falciparum and vivax 6-Oxopurine Phosphoribosyltransferases and Their Prodrugs As Antimalarial Agents.
Keough, D.T., Hockova, D., Janeba, Z., Wang, T.H., Naesens, L., Edstein, M.D., Chavchich, M., Guddat, L.W.(2015) J Med Chem 58: 827-846
- PubMed: 25494538
- DOI: https://doi.org/10.1021/jm501416t
- Primary Citation of Related Structures:
4RAB, 4RAC, 4RAD, 4RAN, 4RAO, 4RAQ - PubMed Abstract:
Hypoxanthine-guanine-[xanthine] phosphoribosyltransferase (HG[X]PRT) is considered an important target for antimalarial chemotherapy as it is the only pathway for the synthesis of the purine nucleoside monophosphates required for DNA/RNA production. Thus, inhibition of this enzyme should result in cessation of replication. The aza-acyclic nucleoside phosphonates (aza-ANPs) are good inhibitors of Plasmodium falciparum HGXPRT (PfHGXPRT), with Ki values as low as 0.08 and 0.01 μM for Plasmodium vivax HGPRT (PvHGPRT). Prodrugs of these aza-ANPs exhibit antimalarial activity against Pf lines with IC50 values (0.8-6.0 μM) and have low cytotoxicity against human cells. Crystal structures of six of these compounds in complex with human HGPRT have been determined. These suggest that the different affinities of these aza-ANPs could be due to the flexibility of the loops surrounding the active site as well as the flexibility of the inhibitors, allowing them to adapt to fit into three binding pockets of the enzyme(s).
Organizational Affiliation:
The School of Chemistry and Molecular Biosciences, The University of Queensland , St. Lucia, Brisbane 4072, Queensland Australia.