Structural basis of IFN receptor recognition by TYK2
Wallweber, H.J.A., Tam, C., Franke, Y., Starovasnik, M.A., Lupardus, P.J.(2014) Nat Struct Mol Biol
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
(2014) Nat Struct Mol Biol
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Non-receptor tyrosine-protein kinase TYK2 | 567 | Homo sapiens | Mutation(s): 0 Gene Names: TYK2 EC: 2.7.10.2 | 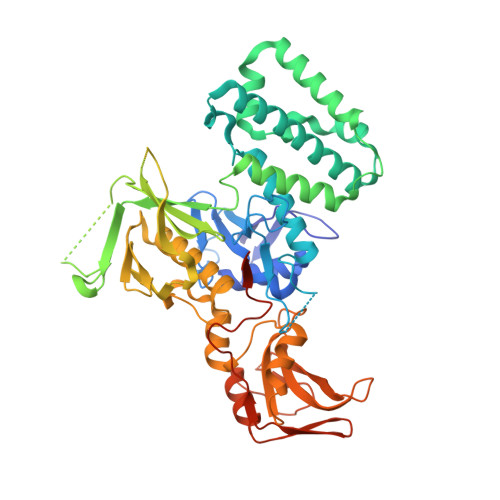 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P29597 (Homo sapiens) Explore P29597 Go to UniProtKB: P29597 | |||||
PHAROS: P29597 GTEx: ENSG00000105397 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P29597 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Interferon alpha/beta receptor 1 | 34 | Homo sapiens | Mutation(s): 0 Gene Names: IFNAR, IFNAR1 |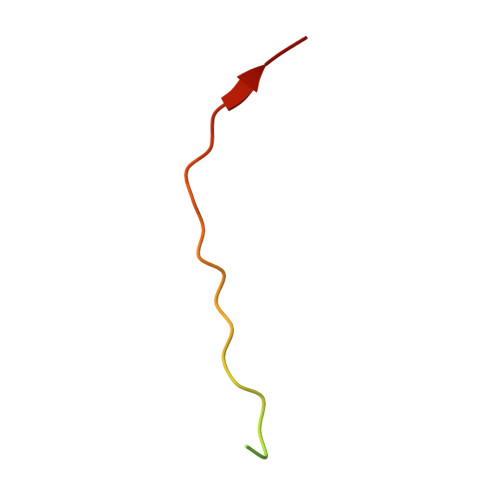  | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P17181 (Homo sapiens) Explore P17181 Go to UniProtKB: P17181 | |||||
PHAROS: P17181 GTEx: ENSG00000142166 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P17181 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GOL Query on GOL | C [auth A], D [auth A], E [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 186.768 | α = 90 |
| b = 52.422 | β = 108.09 |
| c = 70.163 | γ = 90 |
| Software Name | Purpose |
|---|---|
| SSRL | data collection |
| SOLVE | phasing |
| BUSTER | refinement |
| HKL-2000 | data reduction |
| SCALA | data scaling |