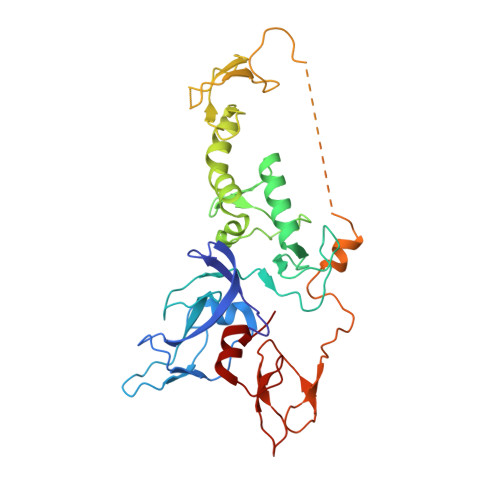
Structure and function of Parkin E3 ubiquitin ligase reveals aspects of RING and HECT ligases.
Riley, B.E., Lougheed, J.C., Callaway, K., Velasquez, M., Brecht, E., Nguyen, L., Shaler, T., Walker, D., Yang, Y., Regnstrom, K., Diep, L., Zhang, Z., Chiou, S., Bova, M., Artis, D.R., Yao, N., Baker, J., Yednock, T., Johnston, J.A.(2013) Nat Commun 4: 1982-1982
- PubMed: 23770887
- DOI: https://doi.org/10.1038/ncomms2982
- Primary Citation of Related Structures:
4I1F, 4I1H - PubMed Abstract:
Parkin is a RING-between-RING E3 ligase that functions in the covalent attachment of ubiquitin to specific substrates, and mutations in Parkin are linked to Parkinson's disease, cancer and mycobacterial infection. The RING-between-RING family of E3 ligases are suggested to function with a canonical RING domain and a catalytic cysteine residue usually restricted to HECT E3 ligases, thus termed 'RING/HECT hybrid' enzymes. Here we present the 1.58 Å structure of Parkin-R0RBR, revealing the fold architecture for the four RING domains, and several unpredicted interfaces. Examination of the Parkin active site suggests a catalytic network consisting of C431 and H433. In cells, mutation of C431 eliminates Parkin-catalysed degradation of mitochondria, and capture of an ubiquitin oxyester confirms C431 as Parkin's cellular active site. Our data confirm that Parkin is a RING/HECT hybrid, and provide the first crystal structure of an RING-between-RING E3 ligase at atomic resolution, providing insight into this disease-related protein.
Organizational Affiliation:
Elan Pharmaceuticals, 180 Oyster Point Boulevard, South San Francisco, California 94080, USA.