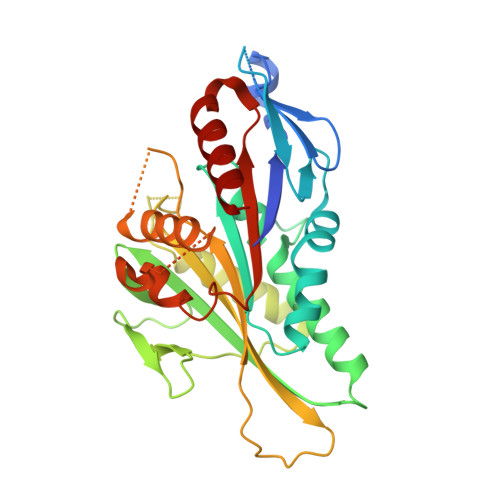
Insight into the molecular mechanism of the multitasking kinesin-8 motor.
Peters, C., Brejc, K., Belmont, L., Bodey, A.J., Lee, Y., Yu, M., Guo, J., Sakowicz, R., Hartman, J., Moores, C.A.(2010) EMBO J 29: 3437-3447
- PubMed: 20818331
- DOI: https://doi.org/10.1038/emboj.2010.220
- Primary Citation of Related Structures:
3LRE - PubMed Abstract:
Members of the kinesin-8 motor class have the remarkable ability to both walk towards microtubule plus-ends and depolymerise these ends on arrival, thereby regulating microtubule length. To analyse how kinesin-8 multitasks, we studied the structure and function of the kinesin-8 motor domain. We determined the first crystal structure of a kinesin-8 and used cryo-electron microscopy to calculate the structure of the microtubule-bound motor. Microtubule-bound kinesin-8 reveals a new conformation compared with the crystal structure, including a bent conformation of the α4 relay helix and ordering of functionally important loops. The kinesin-8 motor domain does not depolymerise stabilised microtubules with ATP but does form tubulin rings in the presence of a non-hydrolysable ATP analogue. This shows that, by collaborating, kinesin-8 motor domain molecules can release tubulin from microtubules, and that they have a similar mechanical effect on microtubule ends as kinesin-13, which enables depolymerisation. Our data reveal aspects of the molecular mechanism of kinesin-8 motors that contribute to their unique dual motile and depolymerising functions, which are adapted to control microtubule length.
Organizational Affiliation:
Institute of Structural and Molecular Biology, Birkbeck College, London, UK.