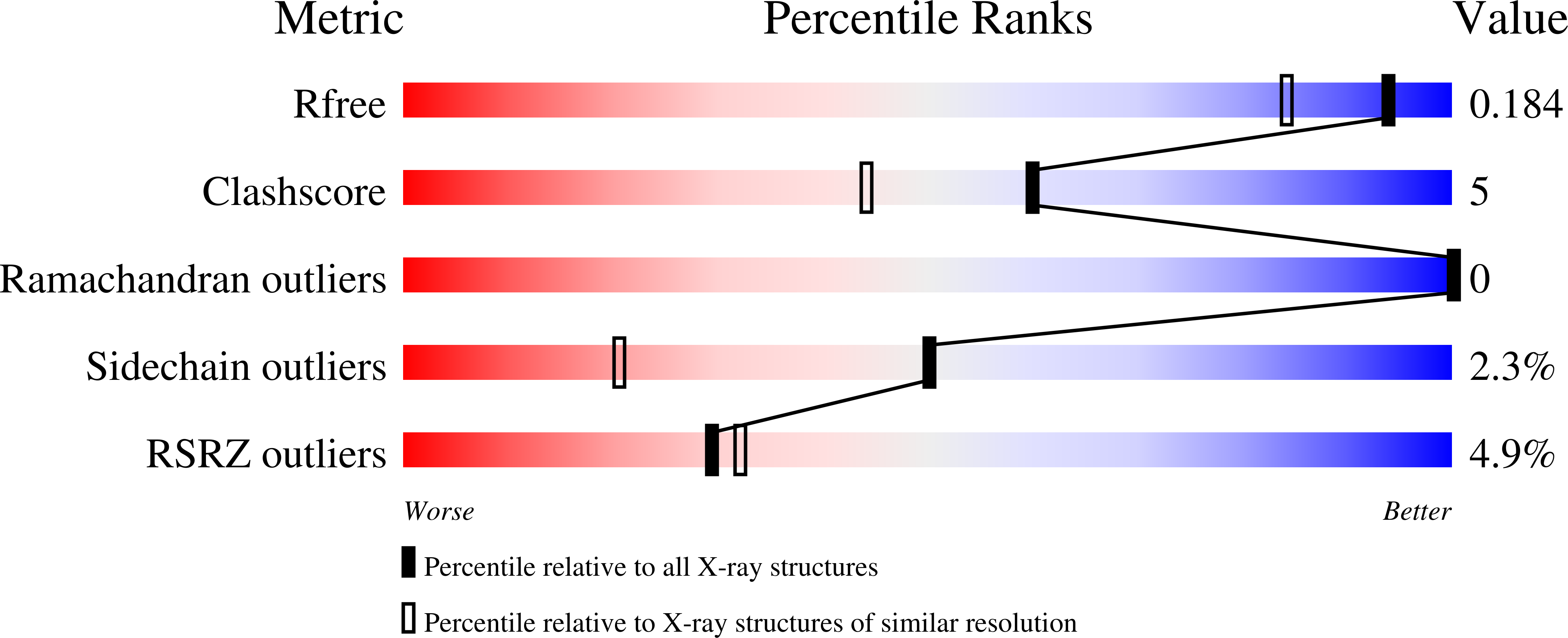
Crystal Structures of Human GluR5 Ligand-Binding Core in Complexes with Novel Marine-Derived Toxins, Dysiherbaine and Neodysiherbaine A, and Their Analogues
Unno, M., Shinohara, M., Takayama, K., Teruya, K., Doh-ura, K., Sakai, R., Sasaki, M., Ikeda-Saito, M.To be published.