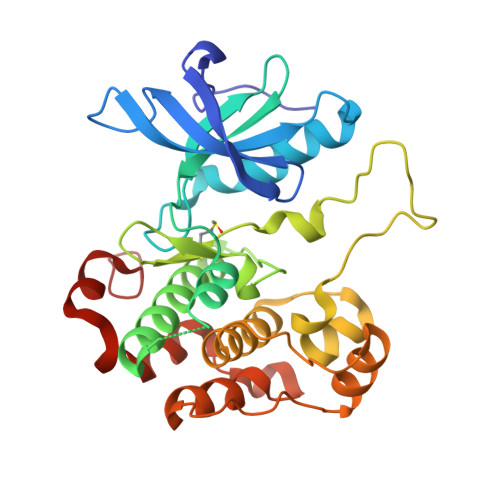
Small-molecule inhibition and activation-loop trans-phosphorylation of the IGF1 receptor
Wu, J., Li, W., Craddock, B.P., Foreman, K.W., Mulvihill, M.J., Ji, Q.S., Miller, W.T., Hubbard, S.R.(2008) EMBO J 27: 1985-1994
- PubMed: 18566589
- DOI: https://doi.org/10.1038/emboj.2008.116
- Primary Citation of Related Structures:
3D94 - PubMed Abstract:
The insulin-like growth factor-1 receptor (IGF1R) is a receptor tyrosine kinase (RTK) that has a critical role in mitogenic signalling during embryogenesis and an antiapoptotic role in the survival and progression of many human tumours. Here, we present the crystal structure of the tyrosine kinase domain of IGF1R (IGF1RK), in its unphosphorylated state, in complex with a novel compound, cis-3-[3-(4-methyl-piperazin-l-yl)-cyclobutyl]-1-(2-phenyl-quinolin-7-yl)-imidazo[1,5-a]pyrazin-8-ylamine (PQIP), which we show is a potent inhibitor of both the unphosphorylated (basal) and phosphorylated (activated) states of the kinase. PQIP interacts with residues in the ATP-binding pocket and in the activation loop, which confers specificity for IGF1RK and the highly related insulin receptor (IR) kinase. In this crystal structure, the IGF1RK active site is occupied by Tyr1135 from the activation loop of an symmetry (two-fold)-related molecule. This dimeric arrangement affords, for the first time, a visualization of the initial trans-phosphorylation event in the activation loop of an RTK, and provides a molecular rationale for a naturally occurring mutation in the activation loop of the IR that causes type II diabetes mellitus.
Organizational Affiliation:
Structural Biology Program, Kimmel Center for Biology and Medicine of the Skirball Institute of Biomolecular Medicine, Department of Pharmacology, New York University School of Medicine, New York, NY 10016, USA.